FY2017 Annual Report

Abstract
The genome contains all the genetic information of a given organism. Decoding the genome therefore provides the molecular basis for understanding every biological phenomenon. Since 2009, the Marine Genomics Unit (MGU) has conducted research in the realm of genome-based biological sciences. By decoding genomes of target marine organisms (mainly invertebrates), we wish to understand comprehensively genetic and developmental mechanisms of marine organisms. The major research fields are (a) evolutionary and developmental genomics of marine invertebrates, (b) environmental genomics of coral reefs, and (c) functional genomics of marine organisms, including algae. We have decoded genome of a coral in 2011, a pearl oyster in 2012, and symbiotic dinoflagellate (Symbiodinium) in 2013, respectively. We further decoded genomes of hemichordates and a brachiopod in 2015, and a brown alga in 2016. In FY2017, in collaboration with Australian researchers, we decoded the genome of the Crown-Thorns-Starfish, a pest of coral reefs. Main results of this year research shall be reported below.
1. Staff
- Professor Noriyuki Satoh
- Group Leaders
- Eiichi Shoguchi
- Staff Scientists
- Keisuke Nakashima
- Konstantin Khalturin
- Ken Maeda
- Takeshi Takeuchi
- Jun Inoue
- Postdoctoral Scholars
- Yuuri Yasuoka
- Koki Nishitsuji
- Yuna Zayasu
- Asuka Arimoto
- Yoshikazu Ohno (JSPS PD Fellow)
- Students
- Kenneth Baughman
- Keita Ikegami
- Yi-Jyun Luo (JSPS DC Fellow)
- Tsai-Ming Lu (JSPS DC Fellow)
- Girish Beedessee (JSPS DC Fellow)
- Yafei Mao (JSPS DC Fellow)
- Kun-Lung Li (Lab rotation student)
- Technical Staffs
- Kanako Hisata
- Sakura Kikuchi
- Seiya Kitanobo
- Research Assistants
- Yuki Yasuoka
- Yoshie Nishitsuji
- Haruhi Narisoko
- Research Administrators
- Ms. Shoko Yamakawa
- Ms. Tomomi Teruya
2. OIST PhD Graduation
We congratulate three students for their OIST PhD Graduation.
3. Research activities and findings
3.1. Environmental genomics
The crown-of-thorns starfish (COTS, the Acanthaster planci species group) is a highly fecund predator of reef-building corals (Fig. 1a). COTS population outbreaks cause substantial loss of coral cover, diminishing the integrity and resilience of reef ecosystems in Indo-Pacific area (Fig. 1c). In collaboration with Australian researchers, we sequenced genomes of COTS from the Great Barrier Reef, Australia and Okinawa, Japan (approximately 380-Mbp genome that contains ~24,500 protein coding genes). We challenged to identify gene products that underlie species-specific communication and could potentially be used in biocontrol strategies. We especially focused on water-borne chemical plumes released from aggregating COTS, which make the normally sedentary starfish become highly active (Fig. 1a, b). Peptide sequences detected in these plumes by mass spectrometry are encoded in the COTS genome and expressed in external tissues. The exoproteome released by aggregating COTS consists largely of signaling factors and hydrolytic enzymes, and includes an expanded and rapidly evolving set of starfish-specific ependymin-related proteins (EPDRs). These secreted proteins may be detected by members of a large family of olfactory-receptor-like G-protein-coupled receptors that are expressed externally, sometimes in a sex-specific manner. Therefore, our study provides insights into COTS-specific communication that may guide the generation of peptide mimetics for use on reefs with COTS outbreaks.
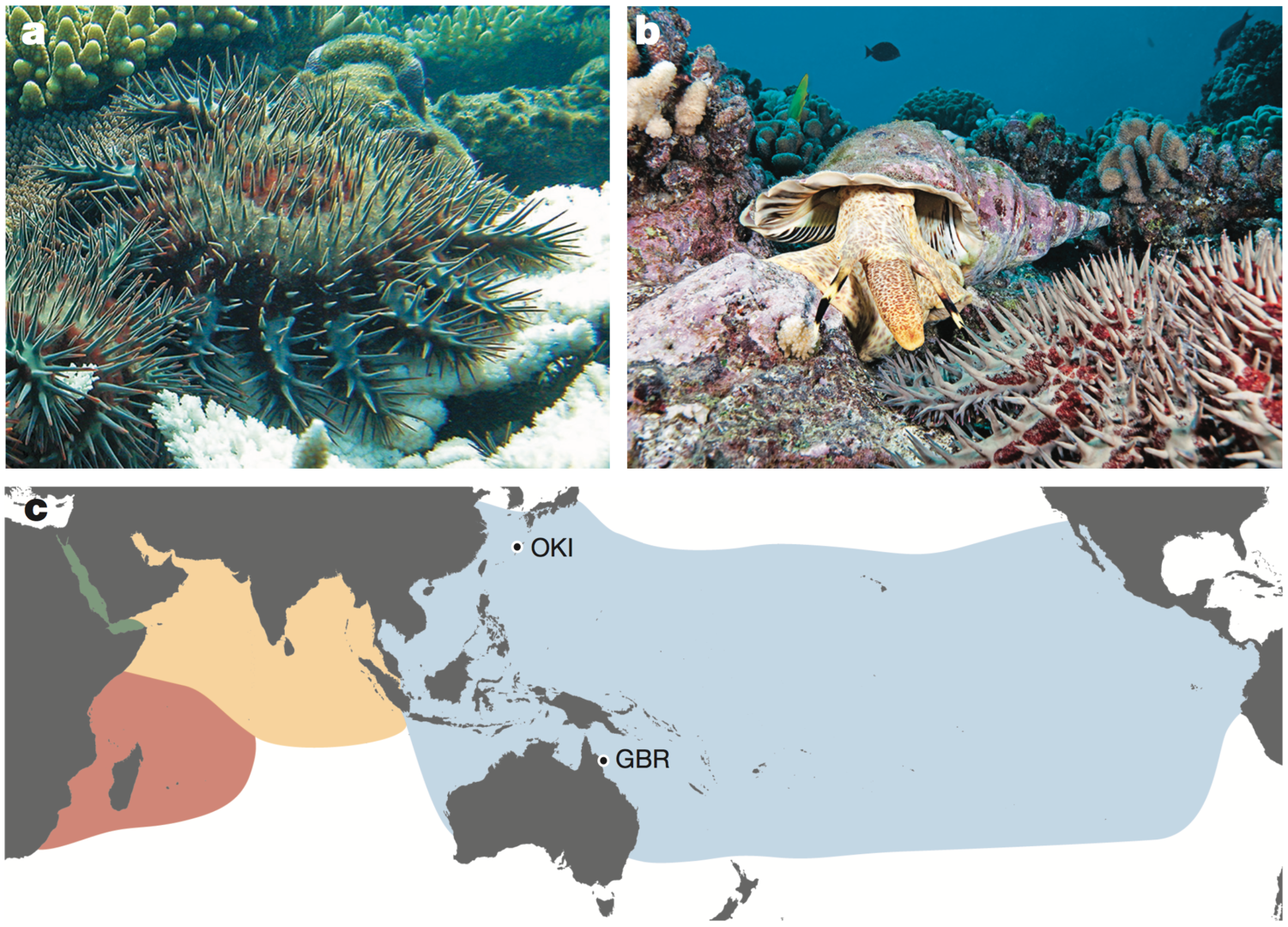

Fig, 1.1. a, Adult COTS predating on coral. White coral skeleton (foreground), unconsumed coral (background). Photo by the Australian Institute of Marine Science. b, A COTS (foreground) and its predator, the giant triton. Photo by Oceanwide Images. c, Global distribution of COTS and the collection sites of the two individuals sequenced. Blue, yellow, pink and green, Pacific Ocean, north Indian Ocean, south Indian Ocean and Red Sea clades, respectively.

Fig. 1.2. a, Tissue expression of the COTS EPDR genes. b, Phylogeny of EPDR proteins. COTS genes are labelled and are marked with red lines; other asteroids, two shades of orange and yellow lines; sea urchins, dark green; hemichordates, light green; molluscs, pink; annelids, purple; cnidarians, black; and vertebrates, blue. The three clades to which COTS sequences belong are indicated by the outer circle. The asterisk denotes the fish-specific true ependymin clade. c, One of the COTS EPDR gene clusters on scaffold 218, with exons (grey bars and arrowheads), intergenic regions and introns (thin black lines) and direction of transcription (arrowhead at end of coding sequence) shown. Scale bar, 10 kb. In all panels, EPDRs secreted by COTS into the seawater are highlighted by red or green.
3.2 Developmental and evolutionary genomics
Of approximately ten publications in the field of Developmental and Evolutionary Genomics this year, we wish to report here a study of phylogenetic position of dicyemids. Dicyemids have long fascinated biologists because of their highly simplified body organization, but their phylogenetic position remains poorly known. We newly sequenced transcriptomes of Dicyema japonicum (Fig. 2.1), which were complemented with published transcriptomic data. Based on comparison of 348 orthologs and 58,124 amino acids, Maximum likelihood and Bayesian inference analyses both placed D. japonicum in a clade with Orthonectida with strong statistical support (Fig. 2.2). Furthermore, maximum likelihood analyses placed the Dicyemida + Orthonectida clade within the Gastrotricha, while in Bayesian inference analyses, this clade is sister group to the clade of Gastrotricha + Platyhelminthes (Fig.2.2). This suggests that dicyemids may share a common ancestor with gastrotrichs and platyhelminths, rather than with mollusks and annelids. Thus our analysis supports the traditional acoeloid–planuloid hypothesis of a nearly microscopic, non-coelomate common ancestor of spiralians.

Fig. 2.1. Morphology of dicyemids. a, Dicyemid embryos (em) develop inside the central axial cell (a, white line), which is covered by a single layer of ciliated epidermal cells. Stained nuclei, plasma membrane, and cilia, showing the simple morphology of a sexually reproductive adult Dicyema japonicum with no coelom, gut, or other organs. Sperm cells and eggs produced by hermaphroditic gonads (g) fertilize, and embryos develop inside the central axial cell. Anterior is to the left. b, The central axial cell is covered with a single layer of ciliated epidermal cells (ep). c, Developing larvae produced sexually, possess long cilia for motility to reach a new host. Fluorescence: yellow, plasma membrane labeled with CellMask Deep Red; blue, nuclei labeled with DAPI; red, acetylated tubulin. Scale bar, 20 μm
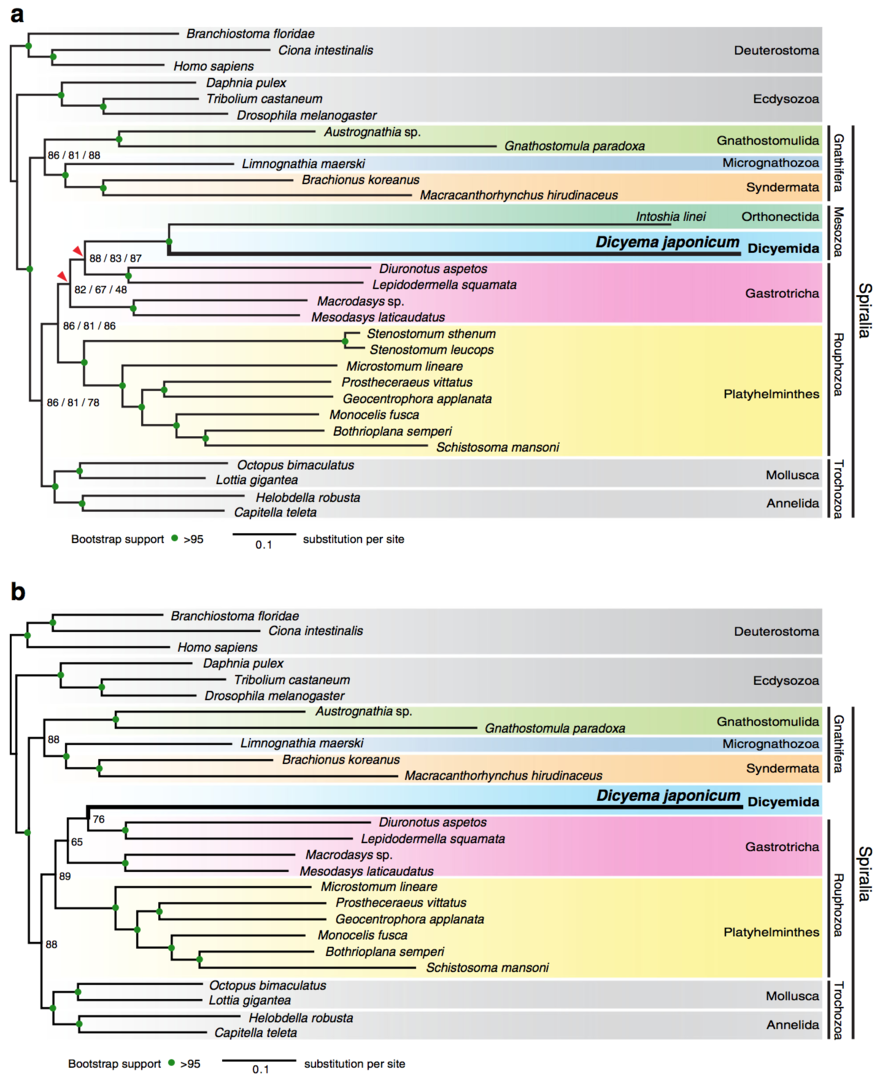 Fig. 2.2. Maximum likelihood analyses suggest that the Dicyemida have a close affinity to the Orthonectida, and are nested within the Gastrotricha. a, The maximum likelihood (ML) tree inferred from a dataset covering 29 taxa, with 348 orthologs, 58,124 amino acids, and 6% missing data. This tree topology is consistent with ML trees from analyses of two sub-datasets filtered to remove systematic biases. Analyses were executed under the GAMMA model of rate heterogeneity with 100 bootstrap replicates using RAxML. The Dicyemida displays close affinity to the Orthonectida, and both are nested within the Gastrotricha. Bootstrap values for three datasets (left to right): Datasets 1–3, respectively. Red triangles indicate different groupings from Bayesian analyses. b, ML tree, inferred from Dataset 3 covering 26 taxa for the taxon-exclusion experiment, indicates that the nesting of D. japonicum within the Gastrotricha probably does not reflect long-branch attraction artifacts. Filled green circles indicate >95% bootstrap support for all datasets
Fig. 2.2. Maximum likelihood analyses suggest that the Dicyemida have a close affinity to the Orthonectida, and are nested within the Gastrotricha. a, The maximum likelihood (ML) tree inferred from a dataset covering 29 taxa, with 348 orthologs, 58,124 amino acids, and 6% missing data. This tree topology is consistent with ML trees from analyses of two sub-datasets filtered to remove systematic biases. Analyses were executed under the GAMMA model of rate heterogeneity with 100 bootstrap replicates using RAxML. The Dicyemida displays close affinity to the Orthonectida, and both are nested within the Gastrotricha. Bootstrap values for three datasets (left to right): Datasets 1–3, respectively. Red triangles indicate different groupings from Bayesian analyses. b, ML tree, inferred from Dataset 3 covering 26 taxa for the taxon-exclusion experiment, indicates that the nesting of D. japonicum within the Gastrotricha probably does not reflect long-branch attraction artifacts. Filled green circles indicate >95% bootstrap support for all datasets
4. Publications
(a) Developmental and Evolutionary Genomics
- Simion, P., Philippe, H., Baurain, D., Jager, M., Richter, D.J., Franco, A.D., Roure, B., Satoh, N., Quéinnec, E., Ereskovsky, A., Lapébie, P., Corre, E., Delsuc, F., King, N., Wörheide, G., Manuel, M. A large and consistent phylogenomic dataset supports sponges as the sister group to all other animals. Current Biology. 27:958-967. doi: 10.1016/j.cub.2017.02.031. (2017)


- Michiue, T., Yamamoto, T., Yasuoka, Y., Goto, T., Ikeda, T., Nagurab, K., Nakayama, T., Taira, M., Kinoshita T. High variability of expression profiles of homeologous genes for Wnt, Hh, Notch, and Hippo signaling pathways in Xenopus laevis. Developmental Biology. 426:270-290 (2017)


- Inoue, J., Yasuoka, Y., Takahashi, H., & Satoh, N. The chordate ancestor possessed a single copy of the Brachyury gene for notochord acquisition. Zoological Letters. 23;3:4. doi: 10.1186/s40851-017-0064-9. (2017)


- Charney, R.M., Forouzmand, E., Cho, J.S., Cheung, J., Paraiso, K.D., Yasuoka, Y., Takahashi, S., Taira, M., Blitz, I.L., Xie, X., Cho, K.W.Y. Foxh1 Occupies cis-Regulatory Modules Prior to Dynamic Transcription Factor Interactions Controlling the Mesendoderm Gene Program. Developmental Cell 40:595-607.e4. (2017)


- Lu, T.M., Kanda, M.,Satoh, N., Furuya, H. The phylogenetic position of dicyemid mesozoans offers insights into spiralian evolution. Zoological Letters 3:6 DOI 10.1186/s40851-017-0068-5 (2017)


- Shimizu, K., Luo, Y.J., Satoh, N., Endo, K.
Possible co-option of engrailed during brachiopod and mollusc shell development.
Biology Letters 13: 20170254. (2017)

- Gyoja, F. Basic Helix-Loop-Helix Transcription Factors in Evolution: Roles in Development of Mesoderm and Neural Tissues. genesis. 55, e23051. (2017)

- Sekigami, Y., Kobayashi, T., Omi, A., Nishitsuji, K., Ikuta, T., Fujiyama, A., Satoh, N., Saiga, H. Hox gene cluster of the ascidian, Halocynthia roretzi, reveals several ancient steps of cluster disintegration during ascidian evolution. Zoological Letters 3:17. (2017)


- Nakano, H., Miyazawa, H., Maeno, A., Shiroishi, T., Kakui, K., Koyanagi, R., Kanda, M., Satoh, N., Omori, A., Kohtsuka, H. A new species of Xenoturbella from the western Pacific Ocean and the evolution of Xenoturbella. BMC Evolutionary Biology. 17:245.

(b) Environmental Genomics
- Hall, M.R., Kocot, K.M., Baughman, K.W., Fernandez-Valverde, S.L., Gauthier, M.E.A., Hatleberg, W.L., Krishnan, A., McDougall, C., Motti, C.A., Shoguchi, E., Wang, T., Xiang, X., Zhao, M., Bose, U., Shinzato, C., Hisata, K., Fujie, M., Kanda, M., Cummins, S.F., Satoh, N., Degnan, S.M., Degnan. B.M.
The crown-of-thorns starfish genome as a guide for biocontrol of this coral reef pest.
Nature, 544:231-234 (2017).

- Ohno Y, Iguchi A, Shinzato C, Inoue M, Suzuki A, Sakai K, Nakamura T.
An aposymbiotic primary coral polyp counteracts acidification by active pH regulation.
Scientific Report 7: 40324. (2017)

- Ponnudurai, R., Kleiner, M., Sayavedra, L., Petersen, J.M., Moche, M., Otto, A., Becher, D., Takeuchi, T., Satoh, N., Dubilier, N., Schweder, T., Markert, S.
Metabolic and physiological interdependencies in the Bathymodiolus azoricus symbiosis.
ISME J. 11:463-477. doi: 10.1038/ismej.2016.124. (2017)

- Campbell, M.A., Nielsen, J.G., Sado, T., Shinzato, C., Kanda, M., Satoh, T.P., Miya, M.
Evolutionary affinities of the unfathomable Parabrotulidae: Molecular data indicate placement of Parabrotula within the family Bythitidae, Ophidiiformes.
Mol. Phylogenet. Evol. 109:337-342. (2017)

- Roberts, R.E., Motti, C.A., Baughman, K.B., Satoh, N., Hall, M. R., Cummins, S.
Identification of putative olfactory G-protein coupled receptors in Crown-of-Thorns starfish, Acanthaster planci.
BMC Genomics. 18:400. (2017)
- Ikeuchi E., Ohno Y., Iguchi A., Nakamura T.
Non-bleached colonies of massive Porites may attract fishes for selective grazing during mass bleaching events.
Peer J. 5:e3470. (2017)
- Nakajima, Y., Wepfer, P.H., Suzuki, S., Zayasu, Y., Shinzato, C., Satoh, N., Mitarai, S.
Microsatellite markers for multiple Pocillopora genetic lineages offer new insights about coral populations.
Scientific Reports 2017; 7: 6729.

- Anbutsu, H., Moriyama, M., Nikoh, N., Hosokawa, T., Futahashi, R., Tanahashi, M.,
Menga, X.Y., Kuriwada, T., Mori, N., Oshima, K., Hattori, M., Fujie, M., Satoh, N., Maeda, T., Shigenobu, S., Koga, R., Fukatsu, T.
Symbiosis for beetle’s hardness: symbiont’s tiny genome for supplying a single amino acid, and host’s control of the synthesis pathway.
PNAS 114: E8382–E8391. (2017)
- Higa, H., Shinzato, C., Zayasu, Y., Nagata, T., Kubo, H.
Restoration efforts for coral reefs by fishery cooperatives ~A case in Onna Village, Okinawa~
J. Japanese Coral Reef Society 19:119–128 (2017) [in Japanese]
(c) Functional Genomics
- Matsuyama, T., Yasuike, M., Fujiwara, A., Nakamura, Y., Takano, T., Takeuchi, T., Satoh, N., Adachi, Y., Tsuchihashi, Y., Aoki, H., Odawara, K., Iwanaga, S., Kurita, J., Kamaishi T., Nakayasu, C.
A Spirochaete is suggested as the causative agent of Akoya oyster disease by metagenomic analysis.
PLoS ONE 12: e0182280. https://doi.org/10.1371/journal.pone.0182280 (2017)

- Takeuchi, T.
Molluscan Genomics: Implications for Biology and Aquaculture.
Curr Mol Bio Rep (2017).
(d) Others (Fish taxonomy and ecology)
- Maeda, K., Tsuhako, Y., Tachihara, K.
Early Development of Mudskippers.
In Fishes Out of Water: Biology and Ecology of Mudskippers. (pp. 69–88). CRC Press, Taylor & Francis Group (2017)
- Maeda, K., Saeki, T., Shinzato, C., Koyanagi, R., Satoh, N.
Review of Schismatogobius (Gobiidae) from Japan, with the description of a new species.
Ichthyological Research 65:56. DOI 10.1007/s10228-017-0593-4 (2017)
- Tran, H.D., Iida, M., Maeda, K.
Downstream migration of newly-hatched ayu (Plecoglossus altivelis) in the Tien Yen River of northern Vietnam.
Environmental Biology of Fishes 100:1329–1341. (2017)
- Kunishima, T., Maeda, K., Tachihara, K.
Newly discovered habitat of a threatened goby, Acanthogobius insularis (Perciformes: Gobiidae), in southern part of Okinawa-jima Island, Japan.
Biogeography 19: 69-74 (2017)



 Yi-Jyun Luo
Yi-Jyun Luo Kenneth Baughman
Kenneth Baughman Keita Ikegami
Keita Ikegami