Marine Genomics Unit (Noriyuki Satoh)

Marine genomics unit
Professor Noriyuki Satoh
norisky at oist.jp

The Earthshot Prize and nomination by PDIE Group
The Prize gives five awards each year for solutions that address global environment problems in five categories: Protect & Restore Nature, Clean Our Air, Revive Our Oceans, Build A Waste-free World, and Fix Our Climate. The PDIE Group stands for “Purpose Driven Innovation Ecosystem” and is one of the top Earthshot Prize nominators. .
We are very proud that “OIST CORAL eDNA” project is nominated by PDIE group for the Earthshot Prize, category of “Revive Our Ocean”.
“The Grand Survey of Coral Reefs in the Pacific Ocean”
(Conservation-Research Alliance for healthy coral reef ecosystems using new eDNA technology)

Coral reefs are the most biodiverse habitats on Earth but are vanishing rapidly due to global environmental change. Conservation groups and communities strive to restore local reefs through coral farming. However, monitoring the diversity and health of reefs is a laborious process, as it relies on the observation by divers. In 2021, OIST’s coral research team developed an environmental DNA (eDNA) technique that enables comprehensive reef monitoring without diving and is more effective for identifying diverse species. The method makes it possible to survey almost all coral species in the world reefs and build a global database for coral conservation.
1. Research Background:
(a) Corals, Symbionts, and their Genomes
Being the most biodiverse marine ecosystems, coral reefs harbor around one-third of all known marine life. Their existence has significant ecological and economic implications for the wellbeing of human society, as coral reefs nurture edible marine species, furnish biochemicals and novel pharmaceutical leads, protect coastlines, support local employment through tourism and recreation, and contribute to regional cultures. The dramatic decline in coral abundance and the wholesale loss of reef habitats due to severe and cumulative disturbances, such as rises of seawater temperature, cyclones/typhoons, and massive outbreaks of crown-of-thorns starfish (COTS), is one of the most pressing environmental issues of our time. As marine managers, communities and governments are calling for the preservation of coral reefs, research in coral biology shall support effective conservation methods by elucidating the molecular mechanisms in corals and their symbionts, especially their stress response and disease resilience at the genomic scale.
OIST Marine Genomics Unit (MGU) is the first team in the world to decode a coral genome, namely Acropora digitifera [Shinzato et al., Nature 476: 320-323 (2011)]. Our research revealed that this coral depends on its symbiont for an amino acid, and therefore the escape of symbionts from the host coral results in severe damages, causing coral bleaching. Since then, in collaboration with the University of Tokyo and others, OIST MGU has decoded 20 species of reef-building corals [Shinzato et al., Mol. Biol. Evo. 38: 16-30, 2020], precisely 77% of decoded coral genomes in the world as of this writing (March 2022).
OIST MGU also succeeded in decoding the genome of a coral-symbiotic dinoflagellate, namely Symbiodinium minutum [Shoguchi et al., Curr. Biol. 23: 1399-1408 (2013)] – a significant endeavor given the large size of this genome, which is a half of the human genome. To date, MGU has decoded three clades of coral-symbiotic dinoflagellates. Thus, OIST MGU is the lab where both coral and dinoflagellate symbiont genomes have been decoded, leading the coral genome research in the world.
Please read on to learn more in detail about our genomic research on corals and coral symbionts.

(b) Environmental DNA (eDNA)
OIST MGU wishes to contribute to worldwide coral reef conservation/preservation using our knowledge of coral genomes. We observe two major challenges in learning about global coral reef health conditions. One is the problem of scale. Coral reefs worldwide suffer from environmental stresses. While some are recovering locally, there is not enough knowledge about the present status of coral reef ecosystems globally. Another challenge is related to coral’s diversity. There are more than 1,300 scleractinian (reef-building) species, 236 genera and 25 families. Conventional coral reef survey methods through diving, or more advanced satellite imaging, are limited in terms of scope and depth of information they can gather. Since 2018, OIST MGU has been developing an alternative survey technology based on environmental DNA (eDNA) metabarcoding. In 2021, we devised successfully primers that can identify 38 genera [Shinzato et al., Front. Marine Sci. 8, 2021], enough to survey almost all coral species commonly found in the tropical and subtropical reefs. Please read on to learn more in detail about our research in coral eDNA metabarcoding.
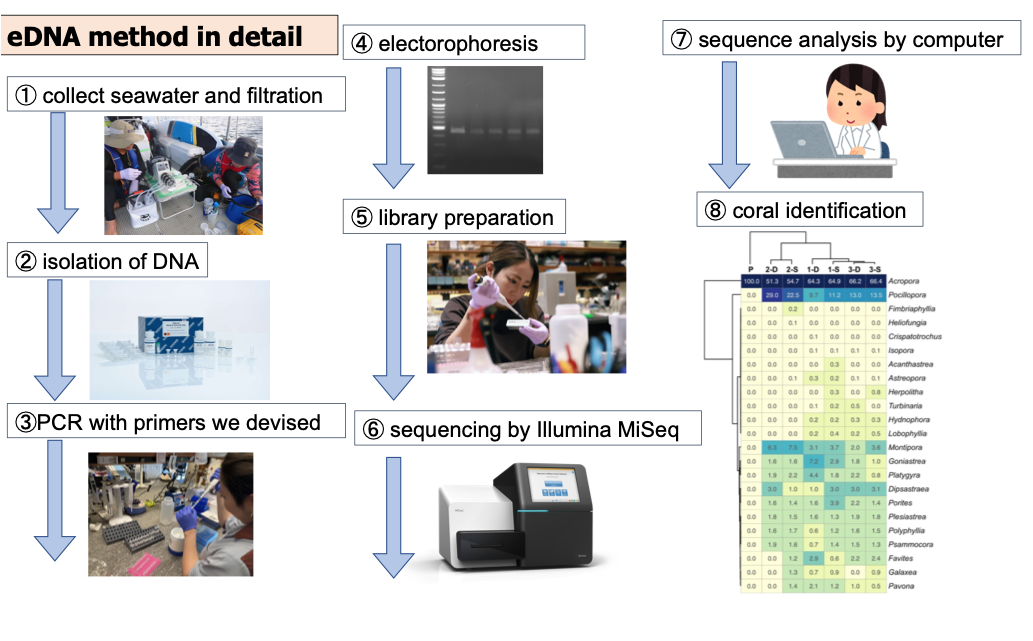
2. OIST CORAL eDNA Project Design: A grand survey of coral reefs in the Pacific Ocean to advance global research and conservation of coral reef ecosystems
OIST MGU proposes to establish an international alliance between conservation and research (CORAL: Conservation Research Alliance) for the preservation of coral reefs. Through this international alliance, we carry out a grand survey of coral reefs in the Pacific Ocean that is enabled by the newly developed eDNA technology. The eDNA information gathered in the survey will be shared with coral reef conservation organizations worldwide through an online database. We envision that such a global knowledge platform of coral reef habitats with genomic information will facilitate greatly the assessments of various local conservation practices, lead to further collaborations, and contribute to innovative solutions for coral reef preservation at the global scale.
a) Partner with organizations such as MANACO Consortium (Front. Marine Sci. 7: 609, 2020) and International Coral Reef Forum (Hainan) for reaching out to researchers and conservation practitioners in the Pacific region.
b) Organize international seminars/workshop to disseminate the new eDNA technology to researchers who will work closely with coral reef conservation organizations in their respective home countries, to expand local coral reef monitoring efforts and to collect eDNA data from major coral reef locations worldwide, with an initial focus on the Pacific Ocean.
c) Establish an eDNA center that will manage all the eDNA information and build a knowledge-sharing platform for coral reef conservation organizations worldwide to use.
3. Related issues at OIST
a) The OIST Environmental DNA Research Center is proposed. The Center will manage all the eDNA information to share the worldwide coral reef conservation organizations.

b) SDGs
The OIST coral eDNA project is a deep commitment to the United Nations’ Sustainable Development Goals (SDGs) that related to oceans, primarily SDG #14: Life Below Water. This project also intersects with other SDGs, including Clean Water and Sanitation (#6), Sustainable Cities and Communities (#11), Climate Action (#13), and Partnerships for the Goals (#17). Onna Village, the home of OIST, has also been named one of Japan’s SDG Cities.

4. Progress Report
a) Publicity
The major national newspaper “Nihon Keizai Shimbun” (Nikkei) reported OIST CORAL eDNA on March 22, 2022.

Koki Nishitsuji, a member of MGU and OIST CORAL eDNA, received an interview by AZoCleantech about the new eDNA solution for coral monitoring on February 4, 2022.

b) Deeper to Sub-Photic coral reefs
Though it depends on the grade of seawater transparency, coral reefs expand from shallow reefs beyond 20m to 50-80m in depth. However, due to limited conventional diving technique, most studies focus coral reefs at 5-20m in depth. Mesophotic coral reefs have not been systematically surveyed and a solution is needed.
The Kerama Islands, just 40 km east of the Okinawa Island, has been designated as a National Park of Japan due to beautiful coral reefs and high transparency of seawater called “Kerama Blue”. In collaboration with NTT Docomo, Co. LTD, which has excellent under seawater robot technique, we start survey of mesophotic coral reefs in Kerama, with a great help of Zamami village. A combinatorial method of eDNA and underwater robot looks very promising and has the great potential to allow OIST CORAL eDNA project survey coral reefs in the sea deeper than ever before.
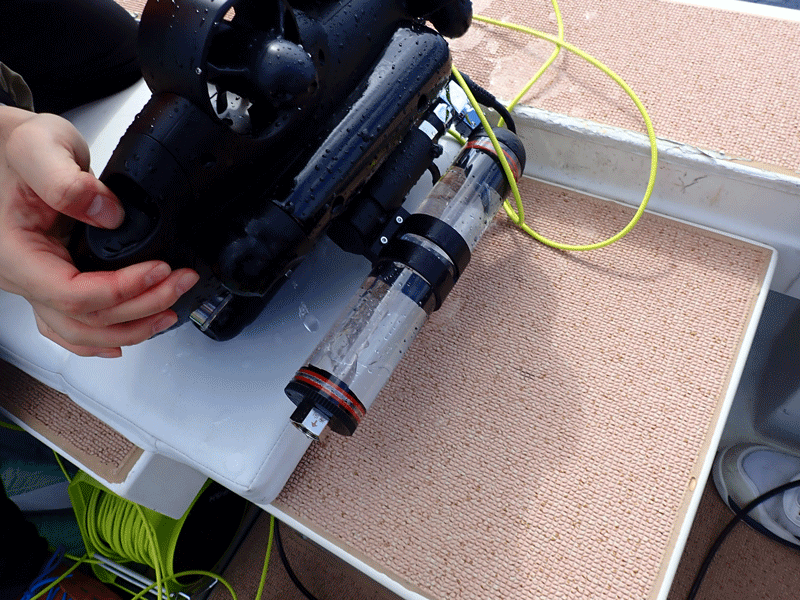
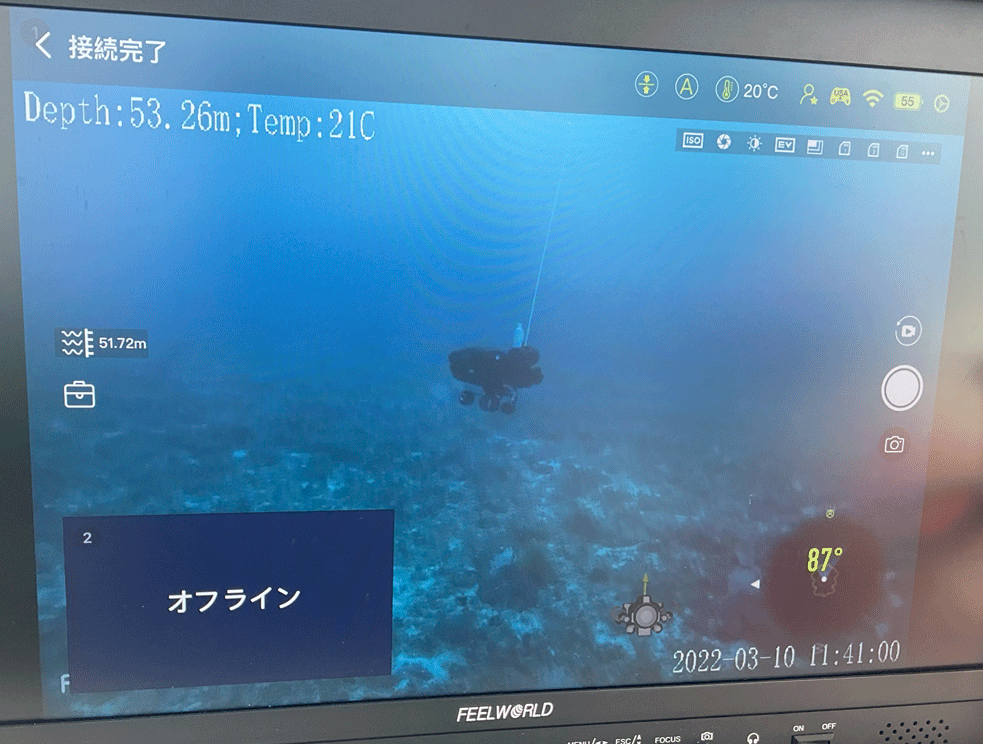

Monitoring robot at 53m in depth Collection of 500mL seawater by robot




