Dynamical Systems Group
Overall goal
A remarkable function of animal's brain is an ability to adapt to environment around them, which is dynamically changed through times. The overall goal of our group is to elucidate a feasible mechanism to realize such a function from the viewpoint of computational rationality. This is so magnificent but too dreamlike, so we set up and tackle with two sub-goals: (1) development of novel learning algorithms and (2) its application to biological networks (e.g. gene, molecular, intracellular cascade.).
Research topics
1. A signal transduction model of synaptic plasticity in cortico-striatum
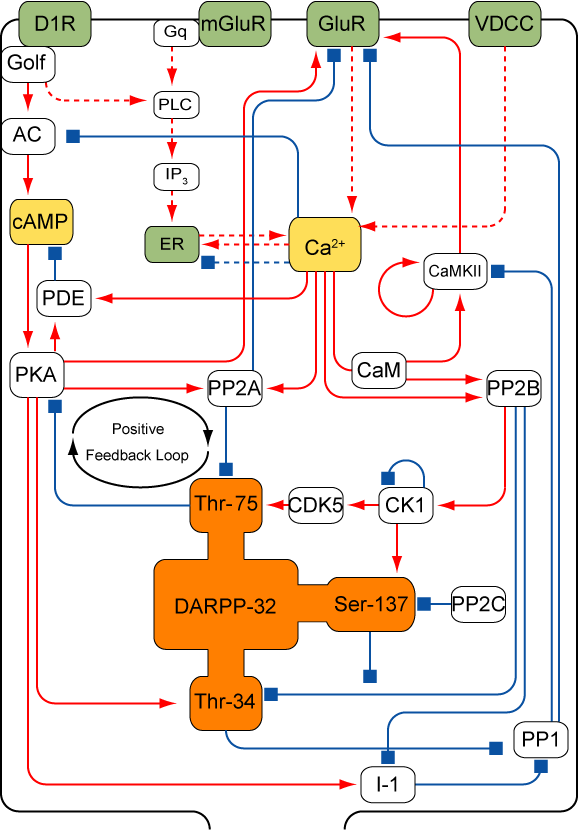 The striatum, the cortical input site of the basal ganglia, receives glutamate from cortex and dopamine from substantia nigra. Recent experimental and theoretical studies suggest that dopamine modulates the synaprtic srength of the cortico-striatum and this dopamine dependent synaptic plasiticity is related with the evaluation of the goodness of the state(reward prediction) in dicision making. Dopamine and glutamate inputs induce the change of synaptic strength by interaction of intracellular molecules. Which intracellular signaling pathway is important for the synaptic plasticity? How do the cascade induce the synaptic plasticity? What is the molecular mechanisms of computation of expected reward? To answer these questions, we construct a model of the intracellular signaling cascade of the striatum based on the biochemical knowledge and analize the kinetic characteristics.
The striatum, the cortical input site of the basal ganglia, receives glutamate from cortex and dopamine from substantia nigra. Recent experimental and theoretical studies suggest that dopamine modulates the synaprtic srength of the cortico-striatum and this dopamine dependent synaptic plasiticity is related with the evaluation of the goodness of the state(reward prediction) in dicision making. Dopamine and glutamate inputs induce the change of synaptic strength by interaction of intracellular molecules. Which intracellular signaling pathway is important for the synaptic plasticity? How do the cascade induce the synaptic plasticity? What is the molecular mechanisms of computation of expected reward? To answer these questions, we construct a model of the intracellular signaling cascade of the striatum based on the biochemical knowledge and analize the kinetic characteristics.
Reference
- Nakano, T., Doi, T., Yoshimoto, J. and Doya, K. (2007). A kinetic model of the molecular cascade for cortico-striatal synaptic potentiation and depression, in The 37th Annual Meeting of the Society for Neuroscience (Neuroscience 2007), BN06, San Diego, CA, 3-7 November, 2007.
2. Bayesian system identification for biochemical reaction systems
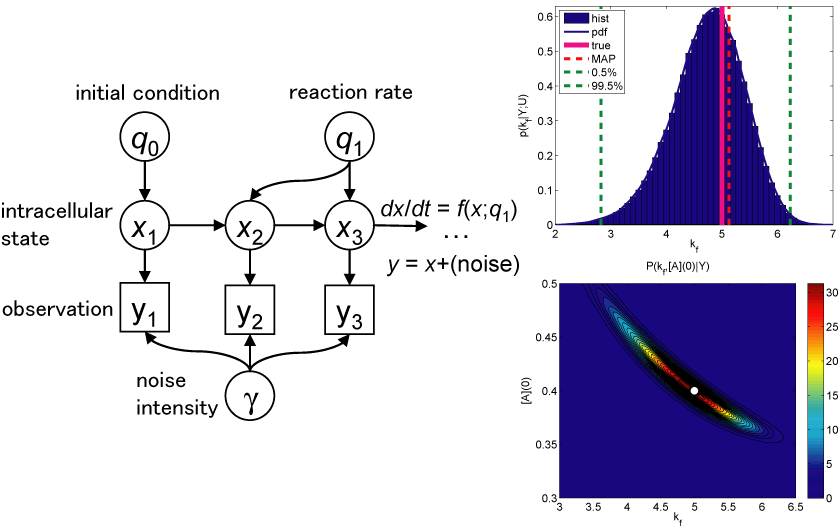
By the recent development in molecular biology, many elementary components making up living organisms - genomes, proteins, neuronal transmitters and so on - have been identified. Then, the trend of the current research in biological science is gradually shifting to dynamic interactions between genes, molecules, and external environments. Mathematical modeling are one good strategy to understand the interactions because the systematic analysis becomes easier by making virtual experiments in various conditions, even if it is infeasible in actual biological experiments. One big issue there is the determination of appropriate parameters and/or structures of the models. In this study, we develop a method to solve this issue based on Bayesian inference. The advantage of this approach is not only to identify the parameters reproducing given data well but also to evaluate the goodness of the model structures simultaneously. Our result will assist in further advance in model-based studies in biology contribute to keeping the consistency between theoretical findings and biological feasibility.
Reference
Yoshimoto, J. and Doya, K. (2007). Bayesian system identification of molecular cascades. in the 14th International Conference on Neural Information Processing, FAA-5, Kita-kyushu, Japan, 13-16 November, 2007.
3. Adaptive state representation for facilitating reinforcement learning in tasks with high-dimensional sensory inputs.
 The ability to make appropriate decisions heavily depends on the brain's flexible ability to represent the physical world depending on the task demand. The precise mechanisms of extracting essential information required for decision making from continuously changing sensory data remain elusive. Recent animal studies indicate that the brain not only extracts statistical regularities of sensory data but also incorporates reward- and action-dependent information in order to construct the representation of the world. In order to realize a highly flexible representation required for appropriate decision making with machine, we aim at developing a reinforcement learning algorithm that automatically adjust the state representation so that it reflects both statistical regularities of raw sensory inputs, and, task demands.
The ability to make appropriate decisions heavily depends on the brain's flexible ability to represent the physical world depending on the task demand. The precise mechanisms of extracting essential information required for decision making from continuously changing sensory data remain elusive. Recent animal studies indicate that the brain not only extracts statistical regularities of sensory data but also incorporates reward- and action-dependent information in order to construct the representation of the world. In order to realize a highly flexible representation required for appropriate decision making with machine, we aim at developing a reinforcement learning algorithm that automatically adjust the state representation so that it reflects both statistical regularities of raw sensory inputs, and, task demands.