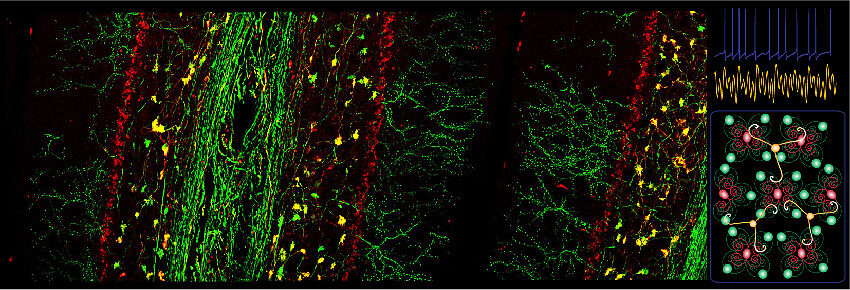
Neuronal Rhythms in Movement Unit (Marylka Yoe Uusisaari)
Our website has been moved to https://www.oist.jp/research/research-units/nrim
Virtually all animal and human behaviour is ultimately enacted through motion, brought about by deliberate and precise muscle activity coordinated across the entire body. Indeed, even simple behaviours and movements (such as raising a glass or sitting down) involve orchestrated adjustments in muscle activities in multitude of kinematic elements of the body. The transformation of abstract and distributed patterns of neural activity in the various planning-related brain structures into these concrete and timely actions requires a reliable temporal code to be used as a framework for behaviour.
The Neuronal Rhythms in Movement (nRIM) unit seeks to uncover this clock and understand its role in spatiotemporal coordination of both motor activity as well as in cognitive functions. The main focus is in the network and dynamics of the olivo-cerebellar system (OCS), believed to be the structure that constructs the temporal framework for behaviour. The mechanisms contributing to this continuous modification of the cerebral computation remain unclear despite decades of investigation. One of the reasons that have hindered advances in this field has been the technical difficulty of examining the relevant elements of the system with a sufficiently wide viewpoint: on one hand, investigating single neurons in the OCS is not likely to lead to understanding of the distributed network dynamics; on the other, methodological constraints have limited examinations of motor behaviour to extremely reduced events of almost singular nature, such as eye blink or whisking.
Recent advances in targeted manipulation of as well as recording from defined neuronal populations, including those residing far from the easily-accessible cortical regions from the brain, are opening new possibilities for dissecting the circuits that generate the neuronal rhythms underlying temporally coordinated movements. The nRIM unit employs methods ranging from genetic manipulations and morphological analysis, in vitro investigation of network dynamics using voltage imaging as well as probing and control of the OCS clock functions in a behaving animal. In order to bring the whole-body kinematics into experimental focus, we will use high-speed recording of the body motions, motion-capture systems and complement the paradigms with virtual/augmented reality environments. The in vivo and in vitro paradigms are bridged by high-resolution, quantitative connectomics and simulation of network-level phenomena.
To see what we have been up to recently, check "Annual reports" in the menu bar on the left!