[Seminar] "Multiscale associative memory recall by modulation of inhibitory circuits" by Dr. Tatsuya Haga
Date
Location
Description
Speaker:
Dr. Tatsuya Haga, Laboratory for Neural Coding and Brain Computing, RIKEN Center for Brain Science
Abstract:
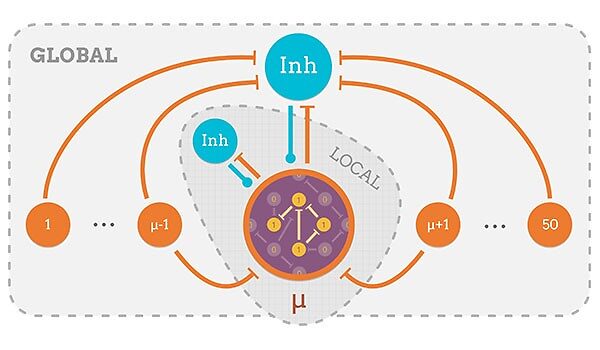
The brain recognizes information in different scales and resolutions. However, how neural networks construct and flexibly process such multiscale information representations has been poorly understood. Here, we demonstrate that a single neural network can offer multiscale information representations based on the graph structure of associative links between memory items. We propose a generalized Hopfield-type associative memory model which can be implemented by excitatory circuits with Hebbian learning between items, and the combination of local and global inhibitory circuits. This network model “analyses” the cluster structure of associative links, and the network flexibly recalls information representation corresponding to each item (microscopic scale), a cluster of items (intermediate scale), and a cluster of interconnected clusters (macroscopic scale) by modulating the ratio between strengths of local and global inhibition. Through mathematical analyses and numerical simulations, we show that recalled activity patterns in this network model correspond to eigenvectors of graph Laplacian, and that modulations of inhibitory circuits control the activation of eigenvectors to change the scale of representations. Eigenvectors of graph Laplacian represent global and local structures in the graph, and they have been used for applications such as optimal graph partitioning and image segmentation. Our model extends these properties to provide a novel framework for multiscale information processing in a single network. Furthermore, our model offers an insight into to the computational role of inhomogeneous neuromodulations of interneuron circuits, which is a focus of recent experimental studies.
Subscribe to the OIST Calendar: Right-click to download, then open in your calendar application.



