FY2021 Annual Report
Developmental Neurobiology Unit
Professor Ichiro Masai

Abstract
Developmental neurobiology unit uses zebrafish as an animal model and will elucidate genetic program that regulate eye development. Specific research projects are to elucidate:
(1) Mechanisms that regulate retinal cell differentiation and neural circuit formation
(2) Mechanisms that regulate photoreceptor degeneration
(3) Mechanisms that regulate lens fiber differentiation
During vertebrate development, the retina is originally derived from anterior neural plate. In this region, six major classes of retinal neurons differentiate and form neural circuits responsible for vision. Thus, the retina provides a good model for studying cell differentiation and neural circuit formation in the developing brain. First, we will investigate mechanisms that regulate cell differentiation and neural circuit formation. Second, we will investigate mechanisms that regulate photoreceptor degeneration. Photoreceptor degeneration is an important topic for medical research, because more than 300 genetic mutations are associated with photoreceptor degeneration in humans. We focus on zebrafish mutants, in which photoreceptor degeneration is caused by defects in protein synthesis in ER, protein transport from ER to the apical photoreceptive membrane region, and phototransduction. Through these mutant analyses, we will determine how photoreceptors monitor abnormalities in cellular functions and trigger apoptosis. We will also investigate the role of microglia in neuronal damage and degeneration in the brain and retina. Third, we will investigate lens development. Especially, we will investigate mechanisms that regulate lens fiber differentiation. Through these projects, we will establish key concepts that govern development of multicellular organisms and also contribute to our understanding of pathological processes of human retinal diseases.
1. Staff
- Dr. Yuko Nishiwaki, Group leader
- Dr. Yuki Sugiyama, Staff scientist (–March 2022)
- Dr. Yuki Takeuchi, Staff scientist
- Dr. Wei-Chieh Chiang, Staff scientist
- Dr. Luis Carretio, Staff scientist
- Ms. Manana Kutsia, phD student
- Ms. Mai Omar Abdulrahman Ahmad, phD student
- Ms. Swathy Babu, phD student
- Mr. Bedish Chatterjee, phD student
- Mr. Hung-Ju Chiang, phD student
- Mr. Dongpeng Hu, phD student
- Ms. Darshini Ravishanker, phD student
- Ms. Sarianna Touvinen, phD student (–August 2021)
- Mr. Akira Kawano, phD student
- Ms. Mariia Pavelchenko, phD student (September 2021–)
- Mr. Yutaka Kojima, Technician
- Dr. Tetsuya Harakuni, Technician
- Dr. Mamoru Fujiwara, Technician
- Mr. Takuya Kamichika (–September 2021)
- Ms. Versha Venkatesha Murthy (–September 2021)
- Ms. Moe Inafuku, Laboratory Assistant
- Ms. Chitose Mizuta, Laboratory Assistant
- Ms. Rui Inoue, Laboratory Assistant
- Ms. Fujino Ishibashi, Laboratory Assistant (–August 2021)
- Ms. Xiujun Huang, Laboratory Assistant (August 2021–)
- Ms. Yuzuki Ikema, Laboratory Assistant (August 2021–)
- Ms. Madoka Makiya, Research Assistant (Tempo staff)
- Ms. Miki Kitamura, Research Assistant (Tempo staff)
- Ms. Mizuki Otake, Research Administrator/Secretary
2. Collaborations
2.1 The Title or Name or Topic of the Collaboration
- Theme: In vivo functional analysis of hypoxia response genes using the zebrafish retina
- Type of collaboration: Joint research agreement
- Researchers: Dr. Ichiro Masai (Developmental neurobiology unit, OIST), Dr. Masayuki Matsushita (Department of Medicine, Ryukyu University)
2.2 The Title or Name or Topic of the Collaboration
- Theme: Role of zebrafish BANP protein in tumor suppression of melanoma
- Type of collaboration: Joint research agreement
- Researchers: Dr. Ichiro Masai (Developmental neurobiology unit, OIST), Dr. Yutaka Kikuchi (Department of Biological Science, Hiroshima University)
3. Activities and Findings
3.1 Mechanism of neural circuit formation in zebrafish retinas
During development, the retina is originally derived from the anterior neural plate. In the retinal region, six major classes of neurons and one types of glial cells differentiate to form the neural circuit, which is responsible for phototransduction and visual processing (Fig. 1). The retinal neural circuit processes visual signals through two synaptic neuropils. At the apical side, the outer plexiform layer (OPL) harbors synapses that transmit input from photoreceptors (PRs) in the outer nuclear layer (ONL) to bipolar (BP) and horizontal cells (HCs) in the inner nuclear layer (INL). At the basal side of the retina, the inner plexiform layer (IPL) is densely packed with synaptic connections formed between BPs and amacrine cells (ACs) in the INL, and retinal ganglion cells (RGCs) in the ganglion cell layer (GCL). The retina contains one type of glial cells called Müller glia (MGs), which span the apico-basal axis of the retina. However, molecular mechanism underlying retinal neural circuit formation remains to be elucidated. In FY2021, we investigated retinal phenotypes of zebrafish strip1 mutants, in which IPL formation is disrupted.
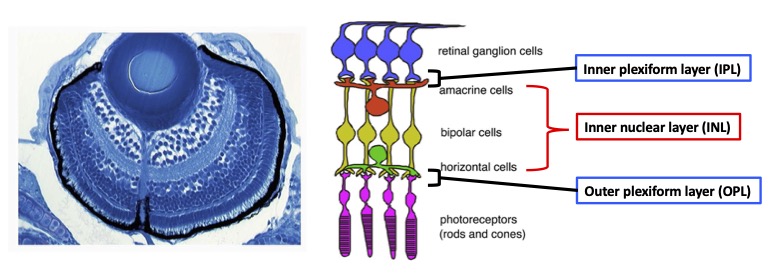
Figure 1: Nerual circuit of vertebrate retina. (Left) Zebrafish retina at 3 dpf. (Right) The retina consists of six major classes of neurons: retinal ganglion cells (RGCs), amacrine cells (ACs), bipolar cells (BPs), horizontal cells (HCs), and two types of photoreceptors (PRs). ACs, BPs, and HCs form inner nuclear layer (INL). Two synaptic layers, inner plexiform layer (IPL) and outer plexiform layer (OPL), are formed at the interface between RGC layer and INL, and between INL and PR layer, respectively.
3.1.1 The role of Strip1 in inner synaptic layer formation
Zebrafish strip1 mutants were identified by zebrafish large-scale mutant screening, which we conducted from 1998 to 2004 (Masai et al., 2003). strip1 mutants shows defects in retinal layer organization, especially IPL (Fig. 2A). IPL is formed by three retinal neurons, RGCs, ACs and BPs. These neurons show an abnormal patterning of dendrites and axons, suggesting that Strip1 is required for proper neurite patterning of these neurons (Fig. 2B). Our cell transplantation analysis revealed that neurite patterning of ACs and BPs is regulated in a cell-non-autonomous manner, indicating that Strip1 does not function in ACs and BPs but rather in RGCs. Indeed, RGCs undergo apoptosis during their maturation in strip1 mutants, which allows ACs and BPs to infiltrate into dying RGC layer (Fig. 2C). This abnormal infiltration of ACs and BPs into RGC layer causes defects in IPL formation. Thus, Strip1 is primarily required for RGC survival.
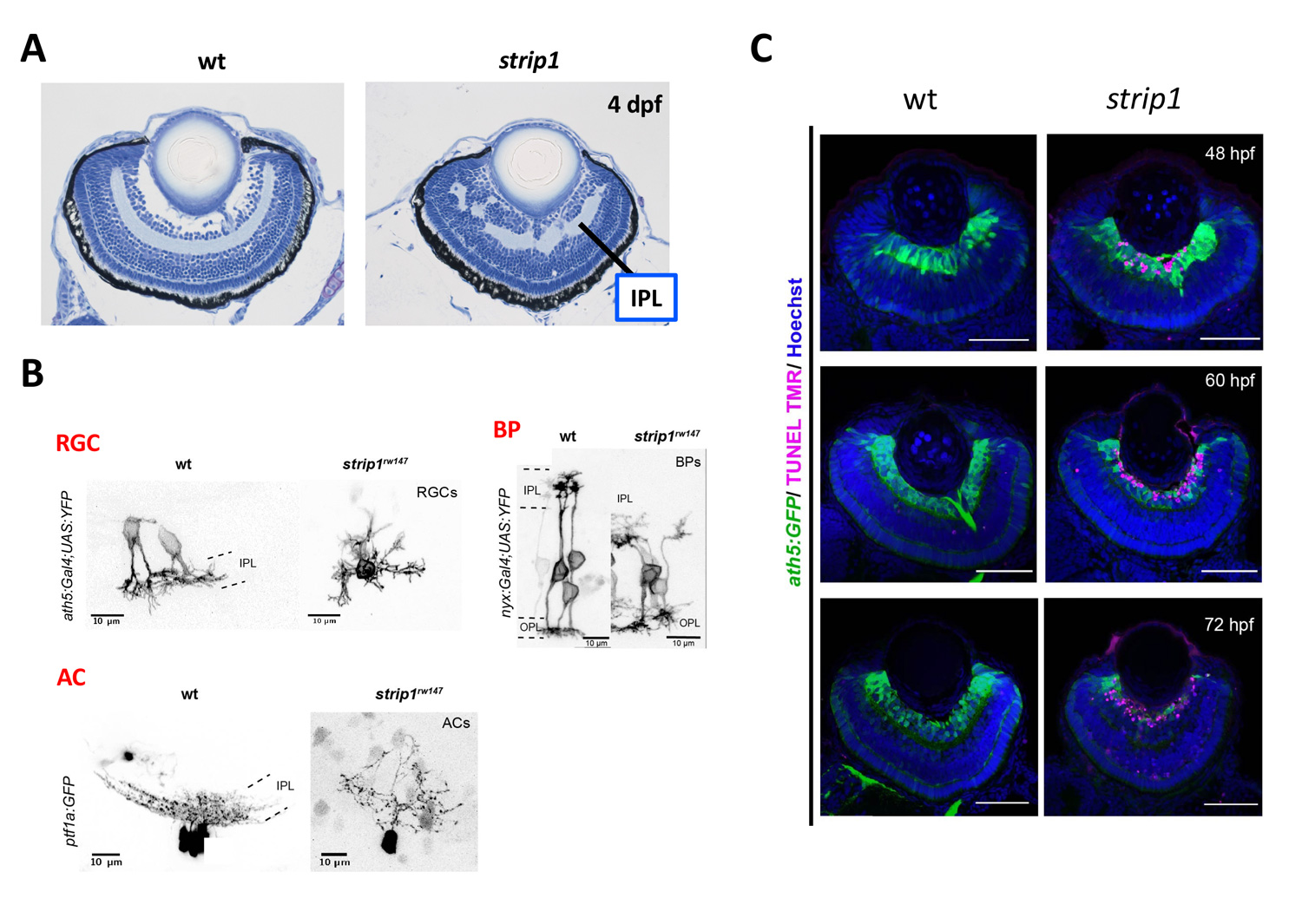
Figure 2: RGCs undergo apoptosis in zebrafish strip1 mutants. (A) Wild-type and strip1 mutant retinas at 4 dpf. IPL formation is affected in strip1 mutants. (B) Neurite morhology of RGCs, ACs, and BPs in wild-type and strip1 mutants. Neurite patterning of RGCs, AC, and BPs is disrupted in strip1 mutants. (C) Visualization of RGCs (green) and apoptosis (magenta) in wild-type and strip1 mutants. In strip1 mutants, apoptosis occurs in RGCs from 48 to 72 hpf.
Strip1 was initially identified as a Striatin interacting protein. Recent studies revealed that Strip1 and Striatin cooperate to form a large protein complex called STRIPAK complex (Fig. 3A). We confirmed that zebrafish Strip1 physically interacts with other component of STRIPAK complex including Striatin3 and that knockdown of Striatin3 shows a similar RGC apoptosis. So, Strip1 and Striatin3 function in RGC survival (Fig. 3B). Our bulk-RNA-seq analysis revealed that the JNK/Jun pathway is abnormally activated in RGCs of strip1 mutants (Fig. 3C). Knockdown of Jun rescues RGC apoptosis in strip1 mutants (Fig. 3D). Thus, STRIPAK complex promotes RGC survival by suppressing JNK/Jun-mediated apoptotic pathway. Since RGC death is associated with optic nerve damage, for example, human glaucoma, we compared RNA expression profile between zebrafish strip1 mutants and zebrafish adult glaucoma model with optic nerve injury. Interestingly, Jun activation is also observed in zebrafish glaucoma model, suggesting that the mechanism underlying Strip1-mediated RGC apoptosis provides an important insight on therapeutic development of human glaucoma.
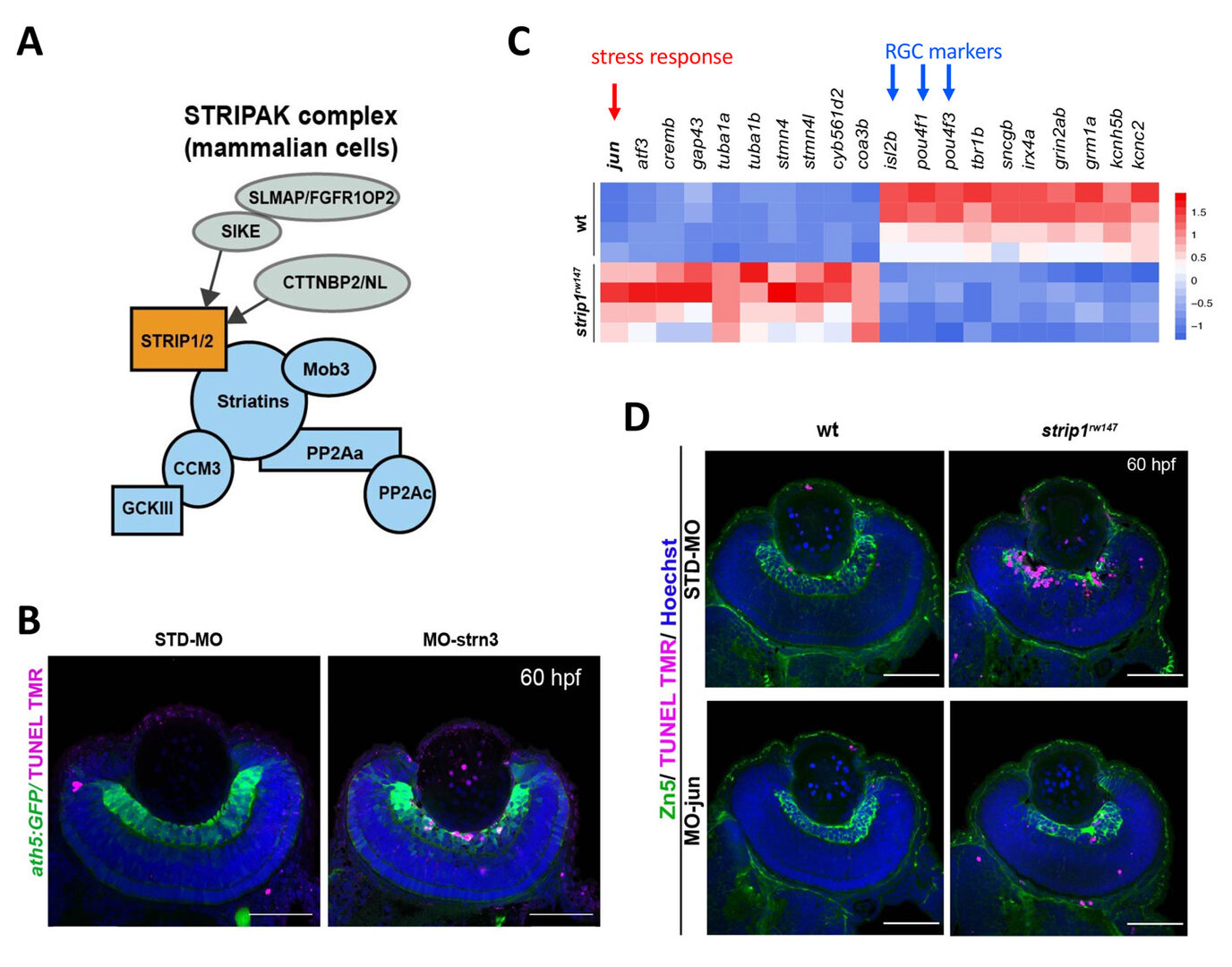
Figure 3: Strip1 adn Striatin3 cooperate to promote RGC survival by suppressing Jun-mediated apoptoptic pathway. (A) Strip1 and Striatin form the STRIPAK complex. (B) Knockdown of Striatin3 causes RGC death as like strip1 mutants. (C) Heat map of upregulated and downregulated genes in strip1 mutants. Top of upregulated gene is jun, whereas top3 of downregulated genes are RGC marker genes. (D) Knockdown of c-Jun inhibits RGC death in strip1 mutants.
Lastly, to elucidate whether IPL formation defects in strip1 mutants depend on only RGC death, we prevented RGC apoptosis in strip1 mutants by overexpression of anti-apoptotic protein, Bcl2. Bcl overexpression rescued RGC death in strip1 mutants (Fig. 4A). We found that AC infiltration into RGC layer is inhibited in Bcl2-overexpressing strip1mutants; however, IPL fails to be formed at the interface between RGCs and ACs but formed at the intermediate position within AC layer (Fig. 4B). We also confirmed that these survived strip1 mutant RGCs show an abnormal patterning of dendrites and axons, indicating that Strip1 is required for proper neurite patterning of RGCs independent of RGC survival, which allows RGCs interacts with ACs to initiate IPL formation program. These data suggest dual roles of Strip1 in IPL formation: (1) RGC survival and (2) RGC neurite patterning (Fig. 5).
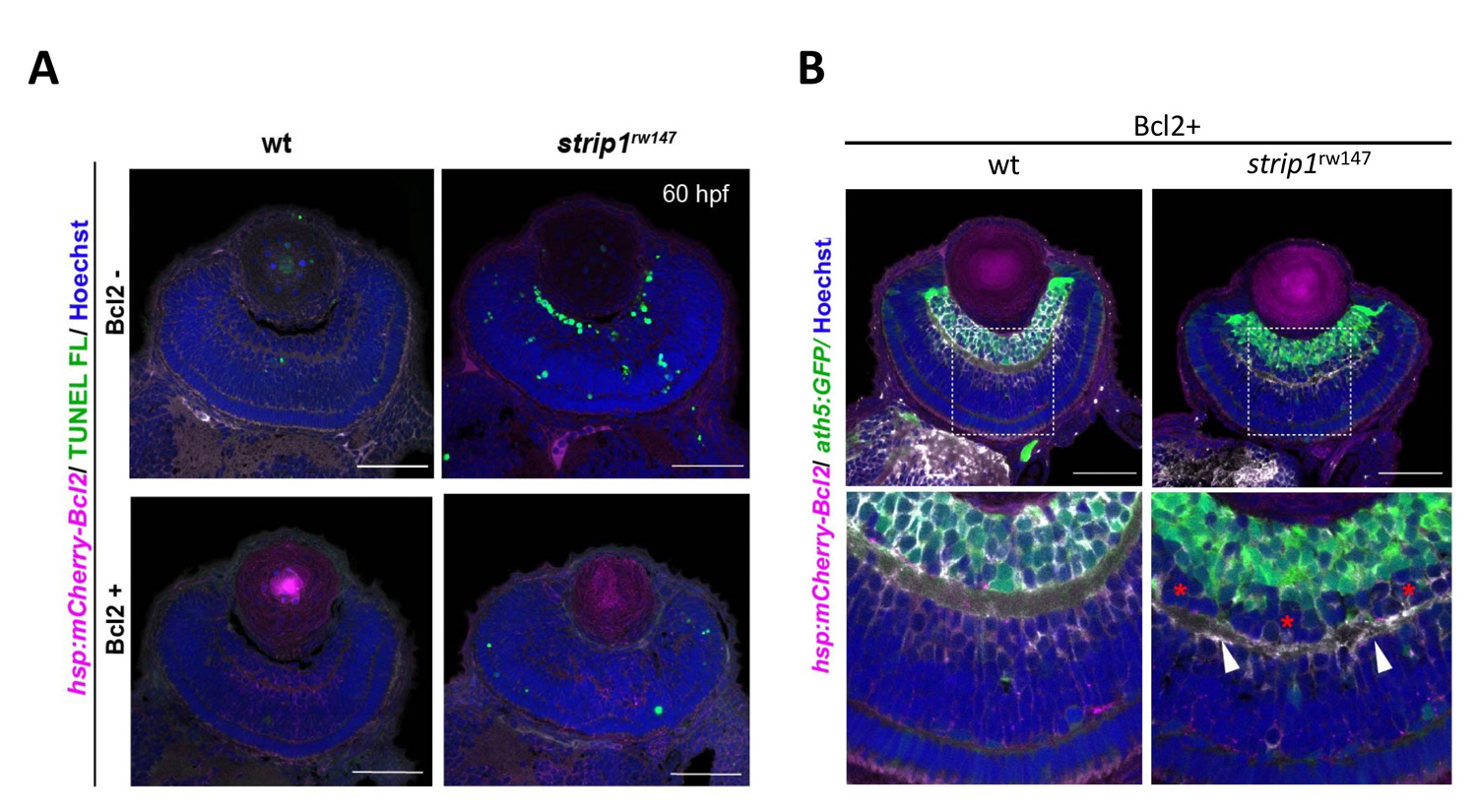
Figure 4: Blockade of RGC death inhibits infiltration of ACs into RGC layer in strip1 mutants, but not IPL formation defects. (A) Bcl2 overexpression prevents RGC apoptosis in strip1 mutants. (B) Bcl2-overexpressing wild-type and strip1 mutant retinas. In strip1 mutants, the interface between RGCs and ACs is maintained, indicating that infiltration of ACs into RGC layer is rescued. However, IPL fail to be formed at the interface between RGCs and ACs, but rather ectopic IPL-like neuropils are observed in the intermediate position of AC layer (arrowheads).
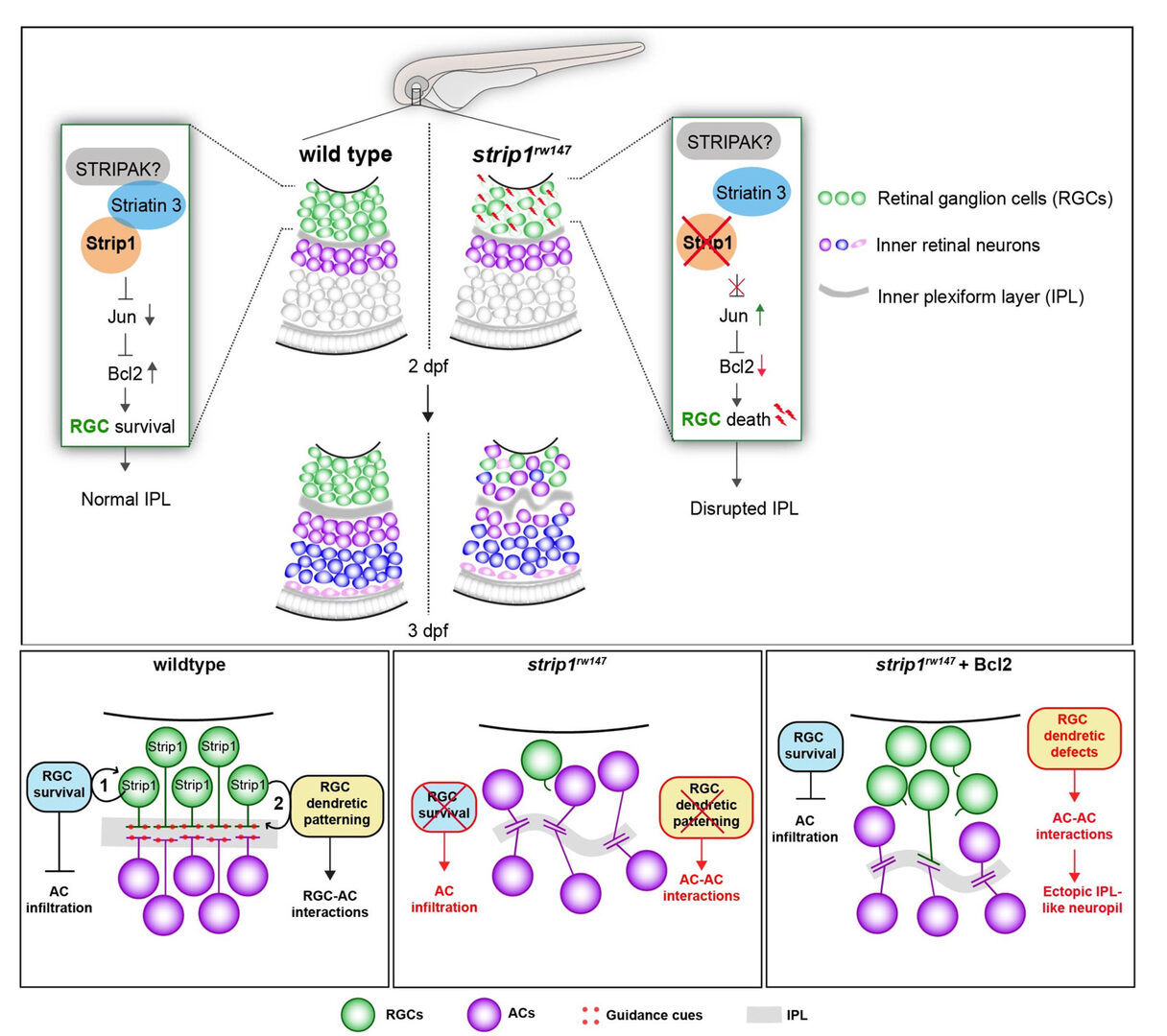
Figure 5: Summary of developmental and molecular events that underlie Strip1 function in inner retinal circuit formation. (A) In wild-type retina, Strip1 suppresses Jun-mediated pro-apoptotic signals through the STIPAK complex, to maintain RGCs during development. In the absence of Strip1, Jun is activated in RGCs leading to severe degeneration of RGCs. Subsequently, cells in the INL abnormally infiltrate the GCL leading to a disrupted IPL. (B) Proposed model for Strip1’s role within RGCs to regulate AC positioning and IPL formation. In wild type, Strip1 regulates (1) RGC survival to prevent AC infiltration, and (2) RGC dendritic patterning to promote RGC-AC interactions. In strip1 mutants, both mechanisms are perturbed, leading to AC infiltration, increased AC-AC interactions, and IPL defects. In Bcl2-rescued strip1 mutants, survived RGCs prevent AC infiltration. However, RGC dendritic defects lead to increased AC-AC interactions and ectopic IPL formation.
3.2 The role of microglia in retinal development and degeneration
Microglia are brain-resident immune cells, originally derived from mesoderm-derivative tissue or the hematopoietic stem cell-lineage, that migrate into brain, and patrol within the brain throughout life. Microglia are thought to eliminate dead or dying neurons to prevent inflammation. Interestingly, microglia-mediated inflammation is required for neuronal regeneration in response to traumatic brain injury in zebrafish (Kyritsis et al., 2012), suggesting more dynamic roles of microglia in neuronal damage. Furthermore, surprisingly, it was reported that microglia facilitate rod cell death in rod-PDE6 mutant mice, which contrasts with our previous view that microglia promote neuronal protection (Zhao et al., 2015). Thus, it is important to understand the role of microglia in photoreceptor degeneration.
3.2.1 Mechanism underlying microglia colonization into the developing retina
In FY2021, we examined normal developmental profiles of microglia and their colonization mechanism to the zebrafish retina. It was reported that microglia are initially generated in lateral plate mesoderm, but that they move to yolk, and enter the brain, including the eyes during zebrafish development (Herbomel et al., 2001). However, it is unknown what kinds of guidance cues enable microglia to move from yolk into developing retina, how microglia are integrated into retinal neural circuits, and also how much cell proliferation contributes to the number of microglia colonizing the retina during development.
To answer these questions, we conducted time-lapse observation using zebrafish mpeg1:GFP or mfap4:tdTomato transgenic lines, which specifically visualizes microglia in zebrafish (Ellett et al., 2011; Walton et al., 2015). We found that microglia progressively enter the optic cup from 24 to 54 hpf through the ventral optic fissure (Fig. 6A). Intraocular blood vessels initially form from the ventral fissure and then expand near the lens, suggesting an overlap with the microglia colonization route. Indeed, microglia are associated with surfaces of intraocular blood vessels. Furthermore, blockade of blood vessel formation reduced the number of microglia in the retina (Fig. 6B), suggesting that blood vessel plays a role in guidance scaffold. We also found that microglia enter the neural retina mainly through the retinal neurogenic area (Fig. 6C). These data suggest that neurogenesis functions as a gate that allows microglia to move into the neural retina. It was reported that brain-derived interleukin 34 (IL34) and its receptor, colony-stimulating factor 1 receptor a (csf1ra) regulate colonization of microglia in zebrafish brain and eye (Wu et al., 2018). We confirmed that IL34 is required for microglia colonization into the retina. Thus, our findings suggest a step-by-step guidance mechanism by which microglia move from yolk into the optic cup and colonize the neural retina (Fig. 6D).
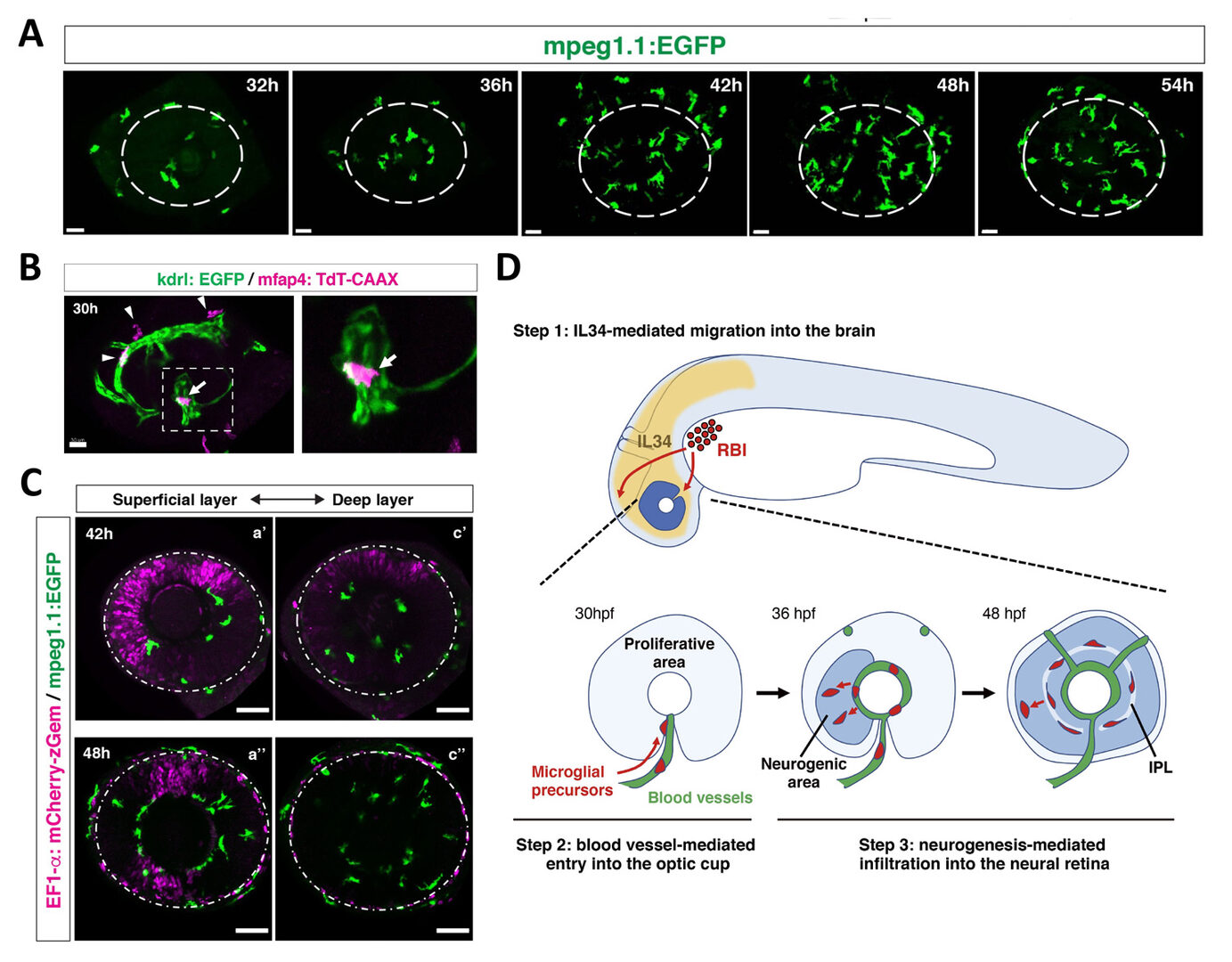
Figure 6: Mechanism of microglial colonization of the developing retina in zebrafish. (A) Three-dimensional confocal images of mpeg1.1:EGFP-positive microglial precursors (green) in the retina from 32 to 54 hpf. Dotted circles indicate the outline of the optic cup. The first microglial precursors appear in the choroid fissure and near the lens at 32 hpf. Microglial precursors in the optic cup progressively increase in number. At 42 hpf, they start to enter retinal tissue and spread into the entire neural retina by 54 hpf. (B) Live confocal images of Tg[kdrl:EGFP; mfap4:tdTomato-CAAX] retinas at 30 hpf. Microglial precursors and blood vessels are visualized using fluorescence of mfap4tdTomato-CAAX (magenta) and kdrl:EGFP (green), respectively. Higher magnification image of a dotted square in the left panel is shown in the right panel. The first microglial precursor (arrow) approaches along developing hyaloid blood vessels near the lens through the choroid fissure. Arrowheads indicate peripheral macrophages outside the optic cup. (C) Live images of Tg[EF1α:mCherry-zGem; mpeg1.1:EGFP] retinas at 42 hpf (upper panels) and 48 hpf (lower panels). Two levels of confocal scanning planes are indicated as superficial (a’, a’’) and deep positions (c’, c’’). mpeg1.1:EGFP positive microglial precursors avoid mCherry-zGem positive proliferating retinal cell area. (D) A possible 3-step model of the guidance mechanism of microglial precursor into zebrafish retina. Step1: IL34 expressed in the brain attracts microglial precursors. Step2: Microglial precursors enter the optic cup along blood vessels. Step3: Microglial precursors infiltrate the neural retina through the neurogenic area.
4. Publications
4.1 Journals (* corresponding authors)
- Ranawat, N., and Masai, I.* (2021) Mechanism underlying microglial colonization of developing neural retina in zebrafish. eLife. 10:e70550. doi:10.7554/eLife.70550.
- Ahmed, M, Kojima, Y., and Masai, I.* (2021) Strip1 regulates retinal ganglion cell survival by suppressing Jun-mediated apoptosis to promote neural circuit formation. eLife. 11:e74650. doi:10.7554/eLife.74650.
4.2 Books and other one-time publications
Nothing to report
4.3 Oral and Poster Presentations
(Oral, International conference)
None
(Poster, international conferences)
- Rodriguez, L. C., Kamichika, T., Izawa, T., and Masai, I. MeCP2 mutant zebrafish shows increases anxiety like behavior, a model for Rett syndrome. (International Zebrafish Conference (IZFC, online), 16-22 June 2021.
(Invited talks)
- Sugiyama, Y. and Masai. FGF controls apico-basal axis rotation and concave to convex conversion of lens fibre cells during spheroidal shape formation of the eye lens (JSDB2021: The 54th Annual Meeting of the Japanese Society of Developmental Biologists, online), 17th – 18th June 2021.
- Nishiwaki, Y., Fujiwara, M, and Masai, I. Dynamin-related GTPase, Drp1, is required for photoreceptor apoptosis in response to vesicular transport defects. (The 73rd Annual Meeting of the Japanese Society of Cell Biology, Kyoto and online) 29 June – 2 July 2021.
(Poster, domestic conferences)
None
5. Intellectual Property Rights and Other Specific Achievements
5.1 The Category or Type of Funding, like External Funding, Awards, etc.
5.1.1 Funding
KAKENHI (grants from the Ministry of Education, Science and Sport/JSPS)
-
PI name: Yuko Nishiwaki
Title: Drp1-mediated formation of ER-mitochondria contact site and its role in apoptisis induction
Category: Grant-in-Aid for Scientific Research (C)
Period: FY2021-FY2023
-
PI name: Yuki Siguyama
Title: Mechanobiological analysis of cell-basement membrane interaction during eye lens morphogenesis
Category: Grant-in-Aid for Scientific Research (C)
Period: FY2021-FY2023
-
PI name: Wei-Chieh Chiang
Title: The mechanism of endoplasmic reticulum proteostasis and proteotoxicity in retinal degeneration.
Category: Grant-in-Aid for Scientific Research (C)
Period: FY2020-FY2023
6. Meetings and Events
6.1 Seminar Title in Full
None
6.2 OIST workshop
None
7. Other
Nothing to report.



