Novel proteomics-based platform for the optimized differentiation of human iPSCs (No. 0122, 0212)
|
|
|
<< Back to all technologies |
Summary
Stem cells are of huge importance to regenerative medicine because of their ability to develop into any specialized cell in the body. Induced pluripotent stem cells (iPSCs) are adult somatic cells which are chemically or genetically reprogrammed to behave like embryonic stem cells. Whilst differentiating iPSCs into basic cell types is relatively easy, huge challenges remain in successfully growing fully functional tissue, such as connected and functioning neurons. For this reason, iPSCs are not yet a therapeutic reality to patients today. A team of OIST researchers led by Prof. Tomoyuki Takahashi has developed a ground-breaking proteomics approach that helps identify both growth and master transcription factors for the differentiation of iPSCs into more functional and better-connected neurons and cardiomyocytes. This represents a major step towards the development of commercial iPSC-based treatments for patients.
Applications
- Regenerative medicine
- Optimized iPSC differentiation
- Diagnostics and patient stratification
Advantages
- Improved differentiation cocktails for healthier, more functional cells
- Identification of novel master transcription factors
- Applicable to any cell type
Technology
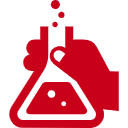
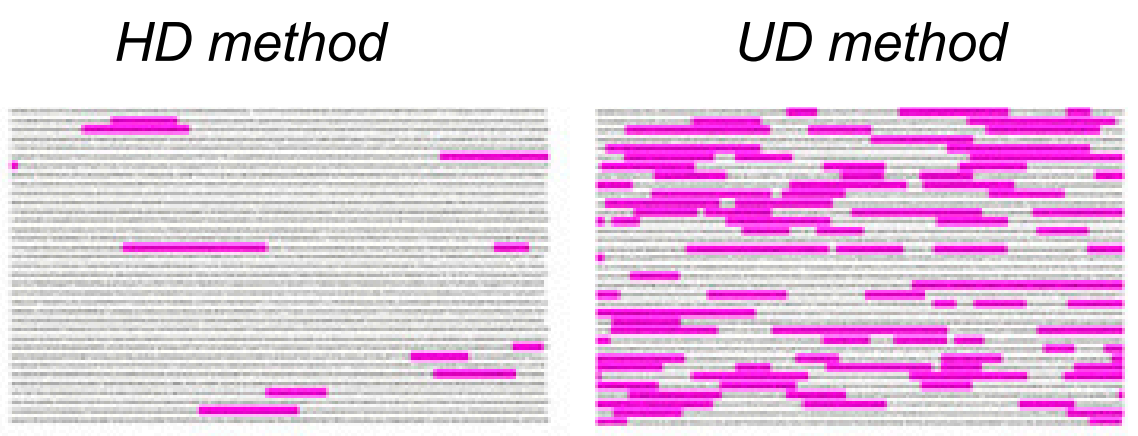
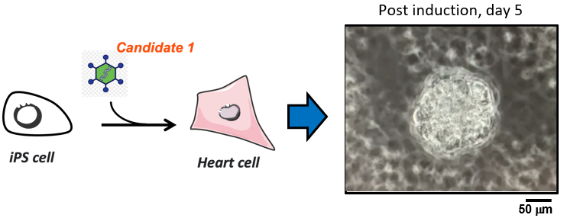
OIST’s innovative ultra-definition (UD) proteomic screening includes a proprietary sample preparation method which allows quantitative identification of three times more receptor proteins than conventional high-definition (HD) proteomics. This enables the optimization of growth factor cocktails and the identification of new master transcription factors for the rapid differentiation of iPSCs into much healthier cells. This UD proteomic screening method can be applied to any type of tissue, such as heart, skin, liver, blood, kidney, lung, retinal, and pancreatic cells. The team is also developing diagnostic and stratification tools through the analysis of the synaptic proteome of mental health patients.
Media Coverage and Presentations
![]() JST Technology Showcase Presentation (JP/EN)
JST Technology Showcase Presentation (JP/EN)
![]() JST Technology Showcase Presentation Slides (JP/EN)
JST Technology Showcase Presentation Slides (JP/EN)
CONTACT FOR MORE INFORMATION
![]() Graham Garner
Graham Garner
Technology Licensing Section
![]() tls@oist.jp
tls@oist.jp
![]() +81(0)98-966-8937
+81(0)98-966-8937