Yoko Nomura, Ph.D.
Science and Technology Associate
Science and Technology Group

About Yoko
I received my Ph.D. in Human Life Science from Japan Women’s University. I first worked as a postdoctoral fellow and a research associate at Research Center for Advanced Science and Technology (RCAST), the University of Tokyo. I subsequently worked as a research associate (Joshu which is equivalent to Jokyo today) at Tokyo University of Technology. In Japan, my main research interest was biosensing for environmental monitoring since my Ph.D.
I moved from Japan to the United States in 2004 and I started my new research projects in the areas of genetic engineering and synthetic biology with Prof. Yohei Yokobayashi in the Department of Biomedical Engineering at University of California, Davis. I planned and executed a wide variety of experiments (cell culture, cloning, RNA work, etc.,) and analyzed the data as an Associate Project Scientist in the Yokobayashi group. I continue to work as a member of Nucleic Acid Chemistry and Engineering (Yokobayashi) Unit here at OIST to work on the projects. I will also start my own research on natural fibers which is related to my primary background in textile science.
My interests- background, current and future projects
1. Design and Engineering of Functional RNAs (as a member of the Yokobayashi Unit)
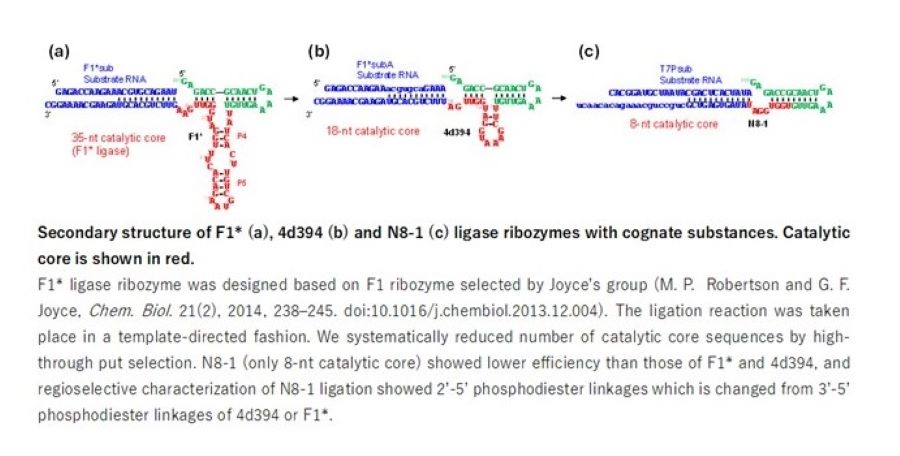
As a member of the Nucleic Acid Chemistry and Engineering Unit, I have been engaged in research to develop synthetic riboswitches for controlling gene expression in E. coli and human cells (HEK293) (K. Fukunaga et al, J. Am. Chem. Soc. 145(14), 2023, 7820–7828. doi: 10.1021/jacs.2c12332. etc.). In addition to this line of research, I recently also started focusing on RNA ligase ribozymes (RNA enzymes) with very small catalytic core sequences which have been largely overlooked in laboratory selection experiments due to their low activities (Y. Nomura and Y. Yokobayashi, Nucleic Acids Res. 47(17), 2019, 8950-8960. doi: 10.1093/nar/gkz729; Y. Nomura and Y. Yokobayashi, Sci. Rep. 13, 2023, 8584. doi: 10.1038/s41598-023-35584-9). Such ribozymes are important when thinking about how RNA sequences were replicated in the primordial “RNA World” which is relevant to the origin of life.
2. Bashofu project (as PI)
Collaborator: Dr. Koji Koizumi (OIST Scientific Imaging Section); Prof. Rie Mori and ex-Specially Appointed Prof. Fumiko Kakihara (Japan Women’s University); Assoc. Prof. Hisako Kawasaki (Niigata University of Pharmacy and Medical and Applied Life Sciences); Bashofu Textile Studio (BTS, Kijoka, Ogimi).
Bashofu is a famous Ryukyuan (old Okinawan) textile which is made from Itobashou banana fibers. The Bashofu of Kijoka is designated as an “Important National Intangible Cultural Property”. However, Bashofu production is declining. To increase productivity, we are characterizing Bashofu and its materials (material plant, fibers etc.) through both scientific and cultural/historical research. Through our efforts, we also aim to contribute to the Okinawan local community.
1) High-quality Bahofu 
Today, high-quality Bashofu is primarily produced as a traditional craft. We have scientifically elucidated the craftsmanship of the traditional method used to make this premium fabric, focusing on how the process alters the microscale morphologies of the materials (Y. Nomura et al., J. Fiber. Sci. Technol. 73(11), 2017, 317-326. doi: 10.2115/fiberst.2017-0039). Building on these results, we collaborated with BTS and others to hold an exhibition and symposium in 2018. Our shared commitment to improving and preserving Bashofu production has fostered a strong partnership with BTS.
We recently proposed a bacterial treatment as a minimal improvement to the traditional method (under review, J. Nat. Fibers). Additionally, we are characterizing the highest quality yarns used for Kimono (Nahagu yarns) from a plant science perspective.
2) Inferior Bashofu
While high-quality Bashofu was produced for the upper class, inferior quality Bashofu was used for summer clothing of ordinary people in Okinawa. We developed a facile fiber extraction method that had been used in inferior Bashofu making (F. Kakihara et al, J. Home Econ. 72(12), 2021, 818-828. doi:10.11428/jhej.72.818).
In cultural history research, we also focused on the issue that Soetsu Yanagi, a prominent figure in the Japanese folk-art movement, did not pay attention to the Bashofu used by ordinary people (R. Mori, F. Kakihara and Y. Nomura, J. Asian Conf. Des. Hist. Theory No. 5, 2024, accepted).
Such research on Bashofu used by ordinary people will be valuable in creating new textiles for daily use, drawing inspiration from the traditional Bashofu craftsmanship.
Please refer to the links for old pubilications: Yoko Nomura: List of Publications



