Eugene Kroll
Research Scientist, OIST, Okinawa, Japan
Research Associate Professor, University of Montana, Missoula MT, USA
Projects
Mechanisms and dynamics of genome changes that occur in populations under stressful conditions.
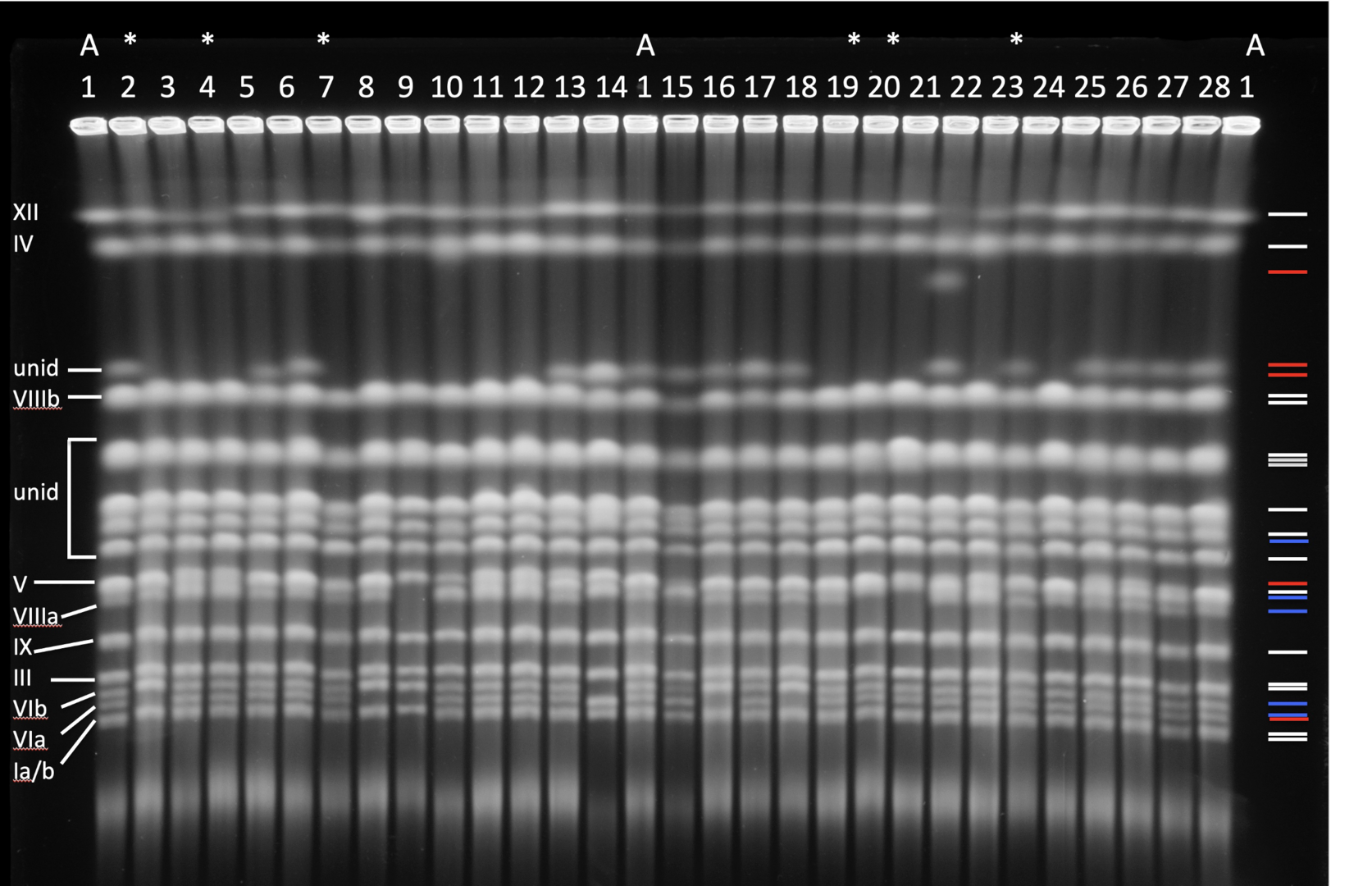
Structural changes in the genome affect adaptive evolution, speciation and tumorigenesis. Genome restructuring is one of the most effective evolutionary tools and genomic restructuring events do accompany and have a causal effect on the aggressiveness of human tumors. Understanding genetic mechanisms that enable populations to adapt to novel environments is required to predict and possibly manipulate evolutionary population fate. To study these mechanisms, we developed an experimental system using the budding yeast Saccharomyces cerevisiae, where starvation serves as a proxy for various environmental conditions that lead to starvation in the wild. We found that starved cell populations rapidly accumulate genomic rearrangements but display only a modest increase in the incidence of point mutations. Survivors with restructured genomes become more resilient to starvation than their common ancestor, and some isolates exhibit reproductive isolation. Because both resilience to starvation and reproductive isolation are strongly associated with genomic restructuring, severe environmental stress may increase the rate of incipient speciation and evolution. The current model we pursue is that external stress might drive large-scale genomic restructuring, which in turn drives significant phenotypic changes in stressed populations. Concurrently, genomic restructuring may cause nascent reproductive isolation that may lead to speciation and thus preservation of newly-acquired adaptive traits. This mechanism, underlain by stress-related genomic restructuring, can explain some macroevolutionary events, such as sudden and large changes in morphology often found in the fossil record. Lastly, tumor cells often accumulate genomic rearrangements at an ever-increasing pace during tumorigenesis. This leads us to ask whether the fundamental genetic mechanisms of such restructuring are shared between yeast and cancer cells.
High-efficiency production of biomolecules: Cell Factories
Molecular biology tools are widely used to create synthetic microorganisms designed to produce high-value biochemicals such as polypeptides (insulin precursors, growth hormones, antimicrobial peptides, antibody fragments), small bioactive molecules (antibiotics, food additives, antimalarials), lipids and other specialty chemicals. However, frequent mutational inactivation of exogenous production genes, particularly those that contain complex heterologous pathways or produce compounds detrimental or toxic to cells, limits production yields, shortens production runtime and increases overall production costs. Additionally, this impedes future designs by imposing high genetic stability as an essential factor in molecular design. We are developing a technology that can halt growth, arrest the cell cycle and thus stabilize the genomes and extend the life cycle of recombinant microorganisms. This cell cycle arrest increases production yields and, over a longer period, may decrease overall production cost. Our technique can be applied to various cells, such as bacterial coliforms, budding and related yeast, molds and metazoan cells. Following our preliminary experiments, we develop a proof-of-concept to show that our technique can stabilize labile genome configurations and increase production yields. If proven successful, this technology should be of interest to companies and academic labs, optimizing engineered microbes to produce high-value biochemicals.
Combination therapy for aggressive tumors
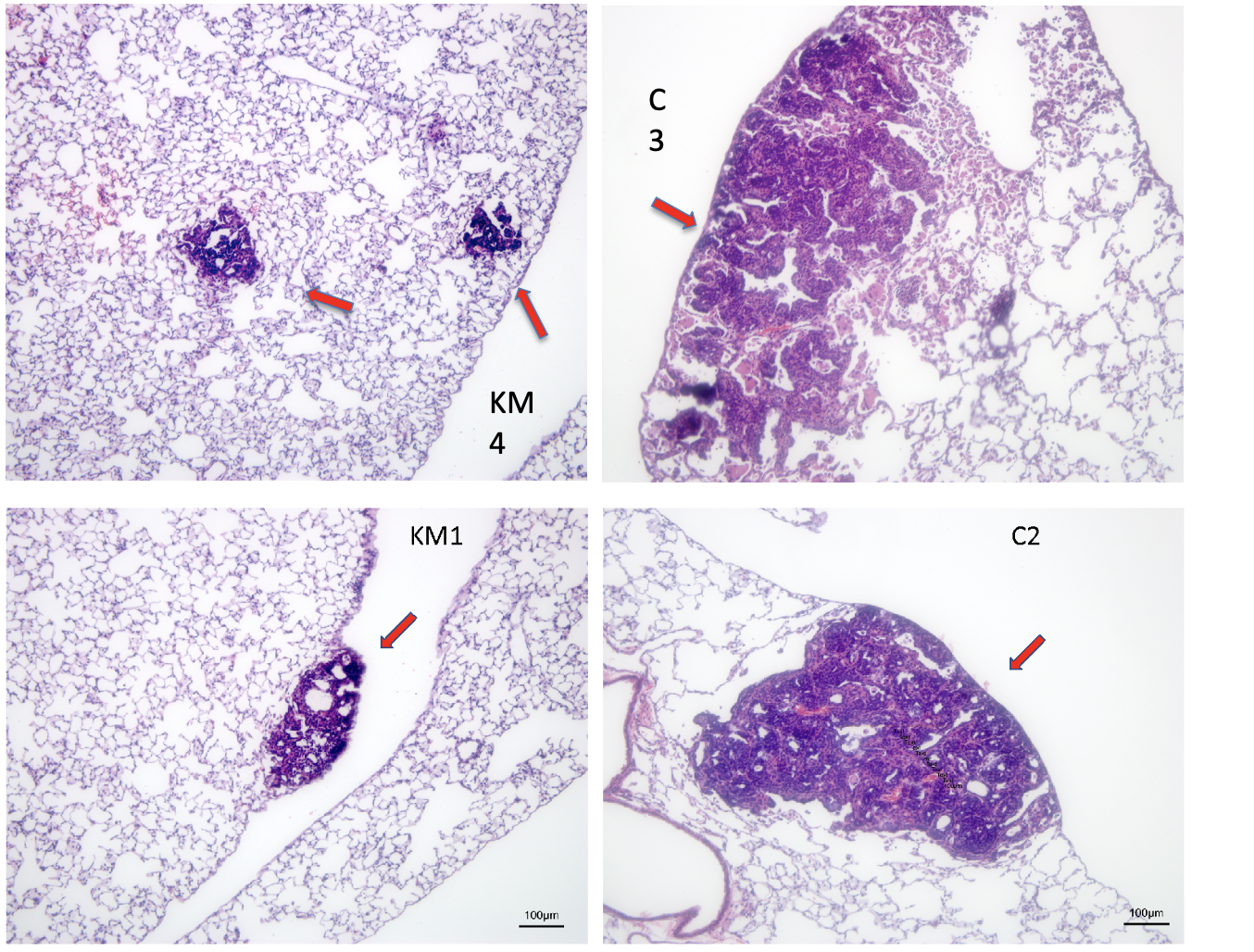
A common feature of aggressively growing tumors and their metastases is intratumoral hypoxia due to insufficient blood supply. Tumor cells adapt to hypoxia by reprogramming their metabolism to rely on glucose fermentation as the primary energetic pathway and drastically increase their glucose intake. This glucose dependence of hypoxic tumor cells can be exploited by limiting the total available glucose. We showed that a combination of a ketogenic diet with clinically relevant doses of metformin significantly reduced blood glucose levels in FVB mice without causing morbidity. Our approach shifts the emphasis of cancer treatment discovery from genetic pathways or protein targets, to which cancer cells tend to evolve and adapt, to a hard-to-escape metabolic vulnerability of cancer by using a combination diet/pharmaceutical therapy approach. While carbohydrate dietary restriction and metformin are well-known interventions, they have yet to be considered in combination for cancer treatment. Significantly, low carbohydrate diets and metformin are compatible in humans outside of the cancer field. If effective, this treatment will be expanded to other hypoxic and glycolytic tumors beyond our current model of breast cancer, particularly those tumors that are refractive to conventional therapies, with the ultimate goal of translating this treatment protocol into human cancer therapy. The combination therapy may also prove to be a significant adjuvant to existing mainline treatments by debulking the hypoxic, typically chemo- and radio-resistant, part of a tumor.



