Human Evolutionary Genomics Unit (Svante Pääbo)
Overview
Until 40,000 years ago we were not the only humans on this planet. Parts of Asia were populated by Denisovans, a group of extinct humans known only from genome sequences, while Neandertals lived in Europe. How are we similar and different to them? In our research unit at OIST, we expand on previous work where we sequenced the genomes of Denisovans and Neandertals by introducing genetic changes that occurred in modern humans and in the extinct hominins into stem cells and mice. We focus on changes that may affect neurodevelopment and behavior, with the goal to identify genetic changes that resulted in phenotypes that differed between modern and extinct humans.
Our Research
The list of genetic variants shared by all or almost all present-day humans but not by the extinct hominins encompasses approximately 30,000 single nucleotide substitutions and about 100 small insertions and deletions. But few of these are likely to influence biological functions. For example, only approximately 100 amino acid substitutions in proteins and about 3,000 substitutions and indels in putative regulatory elements are fixed among humans today and not present in the archaic hominins.
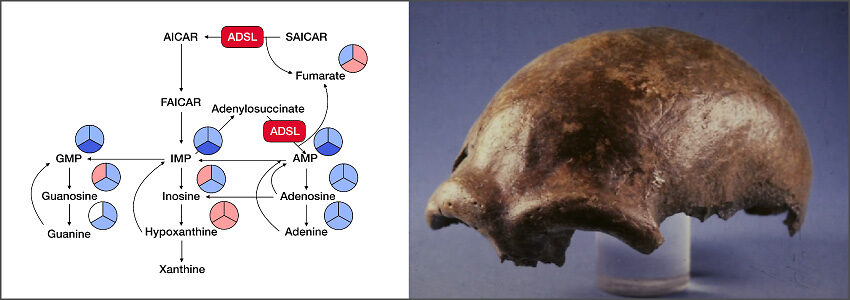
To investigate the physiological consequences of such DNA sequence variants, we have developed novel efficient means to precisely edit multiple nucleotide positions in the genomes of stem cells using CRISPR-Cas9. We use these methods to introduce combinations of changes back to the ancestral, Neandertal-like version in genes that potentially affect related or similar phenotypes. For example, we study genes that have roles in axonal and dendritic outgrowth (e.g. SLITRK1, KATNA1, CALD1), neural progenitor cell division (e.g. KIF18A, CASC5, SPAG5), metabolism and behavior (e.g. ADSL, GLDC, SLITRK1). We also study amino acid substitutions in transcription factors that may affect the expression of many genes. In each case, we introduce multiple substitutions in human embryonic stem cells and analyze the phenotype either in the undifferentiated cells or after differentiation to neurons, astrocytes, or brain organoids. When an effect is detected, we will test the genes, individually and in combinations, to elucidate if variants in a single or multiple genes are responsible. We also introduce some of the variants into mice to test their effects on metabolism and behavior in an organism. For several analyses of both cellular and organismal phenotypes we collaborate with other groups.
A longer term goal is to combine larger numbers of variants to ultimately create human stem cells that are “ancestralized” for certain pathways, certain organelles, or certain functions. Such cell lines will together represent a “library” that can be used by us and others to study cellular traits specific to modern humans or to extinct hominins. For example, we are working towards creating a cell line that carries all, or almost all, of the approximately one hundred amino acid variants that characterize modern humans relative to archaic hominins and thus are “ancestralized” with respect to their proteome. We will also create cell lines “ancestralized” for both amino acid substitutions and regulatory changes in genes encoding transcription factors, in a step towards creating a cells with an “ancestralized transcriptome”.
We also use large cohorts such as UKBioBank and Biobank Japan to investigate if some ancestral variants, or variants specific to Denisovans or Neandertals, occur in low frequency today as this opens up the possibility to study their effects in humans.